Advances in molecular genetic mechanism of meiotic recombination and applications in crop breeding
-
摘要:
减数分裂是真核生物有性生殖产生染色体数目减半的单倍体配子所必需的生命过程。重组是减数分裂的核心事件之一,既增加了同源染色体间遗传信息的交换,又保证了其在减数分裂后期Ⅰ的正确分离。因此,减数分裂重组不仅增加了后代遗传多样性,还是作物遗传育种的基础。通过提高重组频率或改变其分布可以加速农作物育种进程,而降低或抑制重组可以固定杂种优势。近年来对植物减数分裂重组的分子遗传机制的研究取得了很大进展,包括重组的遗传和表观遗传调控机制,重组的遗传操控技术、固定杂交优势和染色体工程等方面。本文针对以上方面进行了全面的总结,这些内容不仅方便了读者对减数分裂重组的理论认知,还拓展了通过调控减数分裂重组操控生物育种的思路。
Abstract:Meiosis is essential for producing haploid gametes during sexual reproduction in most eukaryotes. Homologous recombination is one of the critical events of meiosis prophase I. It not only leads to the reshuffle of genetic information between homologs, but also ensures their proper segregation at anaphase I. Therefore, meiotic recombination is important to facilitate the genetic diversity and evolution among progeny, and also provides the theoretical basis for crop breeding. As expectedly, increasing the frequency of recombination or changing its distribution can benefit crop breeding, while reducing or inhibiting recombination can sustain heterosis. Over the past decades, numerous achievements have been made in understanding and utilizing meiotic recombination in plants, including mechanisms on genetic and epigenetic regulation of meiotic recombination, manipulation technologies on recombination, fixation of heterosis and chromosome engineering. In this review, we summarize the latest findings and technologies for regulating meiotic recombination, which will enable the readers to have an easy access to understand meiotic recombination, and also expand the idea of manipulating breeding through meiotic recombination.
-
水稻、小麦和玉米是全球三大粮食作物,合计约占全球粮食总产量的87%。预计2010—2050年世界对主要粮食作物的需求将增加60%[1]。水稻是单子叶植物的模式植物,也是第1个完成全基因组测序的谷类作物。水稻研究的主要目标是确定每个基因的功能并改善作物的农艺性状[2]。以基因组重测序技术为代表的高通量组学技术在水稻中得到广泛应用,促进了水稻基因功能研究和重要性状遗传改良[3]。
生物信息学是生物科学的一个跨学科分支,可以用来开发用于收集、处理和分析不同生物数据以了解生物功能的方法和工具[4]。在水稻基因组研究中,生物信息学可用于基因序列比对、基因组组装、基因组元件预测和群体遗传学分析等[4-5]。随着越来越多的水稻基因功能得到解析,借鉴水稻生物信息学的研究方法将有助于改善其他作物的农艺性状。此外,生物信息学与前沿基因编辑和合成生物学技术相结合,有望实现对农艺学性状的精准调控和品种的定向改良[3,6]。生物数据正在爆炸性增长,为了有效管理这些大数据,研究者已开发了超过5909个数据库,涉及1525个物种[7]。鉴于水稻生物信息数据库和在线分析工具的激增,对其进行系统的归纳和总结,有利于科研工作者和育种家更好地利用这些资源。
在这篇综述中,笔者系统总结了已报道的水稻相关生物信息学数据库和基于网络服务器开发的在线分析工具,并进行了分类。根据是否只包括水稻一个物种,将数据库分为水稻专门数据库和综合数据库;根据数据集的主要类型和功能,将数据库分为基因组数据库、转录和转录后调控数据库、基因网络数据库和种质资源信息数据库。此外,笔者还总结了可用于基因编辑和智能育种的在线分析工具和数据库。最后,讨论了现有的水稻生物信息数据库和在线分析平台的问题与不足,并对它们在大数据和人工智能时代的发展方向进行了展望。
1. 水稻生物信息数据库
1.1 基因组数据库
高质量的参考基因组信息和准确的基因功能注释,是水稻功能基因组学研究的基础。在21世纪初期,为了推动稻属基因组研究的发展,Wing等[8]提出了OMAP计划(Oryza map alignment project)。在这个框架下,已完成的超过73个稻属种质资源的参考基因组,涉及17个物种,包括所有的二倍体物种和2个异源四倍体(CCDD和KKLL)。由于使用单一参考基因组会导致基因组分析出现严重的偏差,丢失大量群体尺度的遗传变异信息。泛基因组包括一个物种的核心基因和非必须基因,是解决上述问题的有效方法。Yao等[9]基于1483个栽培水稻种质的低覆盖度重测序构建了第1个水稻泛基因组。3KRG (3000 Rice genomes project)完成了3010份国际水稻基因组测序,并获得了它们的线性泛基因组[10]。Zhao等[11]基于67份亚洲栽培稻和普通野生稻高深度二代测序数据,组装了亚洲栽培稻−普通野生稻的线性泛基因组。Qin等[12]构建了33个水稻品种的图形泛基因组。Shang等[13]构建了包括亚洲栽培稻、普通野生稻、非洲栽培稻和短舌野生稻的稻属超级泛基因组。RGI (Rice gene index) 基于同源基因簇构建了水稻泛基因组数据库,提供了丰富的模块和工具,支持对不同种质基因以及基因同源关系进行查询、分析和可视化[14]。Wang等[15]构建了413份国际籼稻的图形泛基因组。此外,已完成了超过6000份水稻种质资源的基因组重测序,这些原始数据大部分储存在SRA (Sequence read archive)和GSA (Genome sequence archive)数据库。针对上述水稻参考基因组、泛基因组及海量的基因组重测序数据,已开发了多个数据库(表1)。根据是否只包含水稻的数据集,这些数据库可以分为综合数据库和水稻专门数据库。NCBI[16]、Ensembl[17]和Phytozome[18]是比较著名的综合数据库,它们提供了大部分已测序物种的参考基因组和比较详细的基因组注释信息;RAP-DB[19]和MSU-RGAP[20]数据库是2个著名的水稻专门数据库,为第1个水稻参考基因组‘日本晴’提供基因组注释资源;基于籼稻参考基因组‘珍汕97’和‘明恢63’,Song等[21]开发了RIGW数据库,包括了基因组学、转录组学和蛋白质−蛋白质相互作用数据集;IC4R提供了一个专门用于整合水稻基因组序列、基因注释和表达水平信息的平台[22];RPAN[10]和Rice-PanGenome[11]是2个为水稻泛基因组分析提供资源和工具的数据库;Rice Genome Hub提供了稻属10个物种的参考基因组信息,并提供了基因组浏览、基因检索和共线性分析等在线工具[23];OryzaGenome[24]和RiceRelativesGD[25]包含了水稻近缘物种的基因和种质资源信息;funRiceGenes[26]是一个专门提供水稻中已克隆基因信息的数据库。综上所述,基因组数据库是水稻所有数据库的基础,也是资源最丰富和内容最完整的一类数据库。然而,由于数据可视化和信息检索方式的差异,容易给用户的使用造成困扰。因此,需要开发一个整合了所有的参考基因组和基因组重测序资源的水稻基因组综合数据库,从而更加规范管理和使用现有的数据资源。
表 1 已发表的水稻基因组数据库Table 1. The published rice genomic databases数据库
Database描述
Description参考文献
ReferenceNCBI 综合数据库、稻属16个物种参考基因组、基因组重测序数据,https://www.ncbi.nlm.nih.gov/ [16] Ensembl 综合数据库、稻属10个物种参考基因组、基因组注释,http://plants.ensembl.org/ [17] Phytozome 综合数据库、‘日本晴’和‘Kitaake’参考基因组、基因组注释,https://phytozome-next.jgi.doe.gov/ [18] RAP-DB ‘日本晴’、IRGSP-1.0参考基因组、基因组注释,http://rapdb.dna.affrc.go.jp/ [19] MSU-RGAP ‘日本晴’、MSU7.0参考基因组、基因组注释,http://rice.uga.edu/ [20] RIGW ‘珍汕97’和‘明恢63’参考基因组、多组学数据、互作数据,http://rice.hzau.edu.cn/rice_rs3/ [21] IC4R 参考基因组、基因组注释、基因表达谱,http://ic4r.org/ [22] Rice Genome Hub 稻属10个物种的参考基因组(32个基因组信息),https://rice-genome-hub.southgreen.fr/ [23] RPAN 3KRG线性泛基因组、泛基因组浏览器,http://cgm.sjtu.edu.cn/3kricedb/ [10] RicePanGenome 线性泛基因组、基因组变异、67个参考基因组,http://db.ncgr.ac.cn/RicePanGenome/ [11] RiceRc 图形泛基因组、33个参考基因组,http://ricerc.sicau.edu.cn/ [12] RiceSuperPIRdb 图形泛基因组、251个参考基因组,http://www.ricesuperpir.com/ [13] RGI 基于同源基因簇的水稻泛基因组、16个水稻参考基因组,https://riceome.hzau.edu.cn [14] OryzaGenome 稻属参考基因组,208个种质基因组信息,涉及19个野生稻和2个栽培稻物种,http://viewer.shigen.info/oryzagenome2detail/ [24] RiceRelativesGD 水稻17个近缘物种基因组和单倍型信息,http://ibi.zju.edu.cn/ricerelativesgd/ [25] funRiceGenes 基因功能数据库、IRGSP-1.0和MSU7.0基因注释,http://funricegenes.ncpgr.cn/ [26] 1.2 转录和转录后调控数据库
通过微阵列或转录组测序,获得不同组织或细胞在各个发育阶段和处理下的基因表达信息,加深了对基因时空表达模型的理解,有利于基因功能的研究[27]。研究者们利用统一的标准流程,将海量的基因表达数据整合在一起,构建了一系列水稻基因表达数据库,包括RiceXPro、CREP、RED、TENOR、eRice和RiceENCODE等(表2)。RiceXPro是基于微阵列表达数据开发的基因转录数据库,提供不同组织或细胞在各个生长发育阶段、逆境胁迫及植物激素处理条件下的基因表达信息[27];CREP包含了‘珍汕97’和‘明恢63’不同组织的微阵列数据集 (Microarray data),支持通过基因序列、基因名称或探针标识来检索基因表达信息[28];RED整合了水稻9个组织的mRNA-seq数据,提供了不同组织和胁迫条件下的基因表达谱[29];TENOR包含了‘日本晴’在不同环境胁迫和激素处理条件下的mRNA-seq数据集[30];PPRD收集了2021年之前发表的11726个水稻mRNA-seq数据集,使用统一流程和最新的参考基因组进行分析和整合[31];eRice包含了‘日本晴’和‘9311’的mRNA-seq数据集,并整合了DNA甲基化和组蛋白修饰数据集[32];RiceENCODE整合了包括ChIA-PET、Hi-C和mRNA-seq等在内的972个数据集,用于检索调控RNA转录的DNA修饰、组蛋白修饰和染色质构象等表观调控[33]。非编码RNA (ncRNA,包括miRNA、lncRNA、siRNA、circRNA等)参与对基因的转录后调控,从而影响植物的生长发育和对环境的响应过程。水稻专门非编码RNA数据库有RiceNCexp、ARMOUR、RiceLncPedia和RiceATM(表2)。RiceNCexp基于mRNA-seq和small RNA-seq (sRNA-seq),对基因和sRNA的转录水平和共表达网络进行了分析,并为用户提供了个性化分析工具[34];ARMOUR整合了7个水稻品种在不同发育时期、组织和胁迫下的miRNA和它们的靶标基因信息[35];RiceLncPedia基于2312个共公数据库获取的RNA-seq数据集,提供了水稻lncRNAs的表达谱、变异位点、lncRNA之间和lncRNA与编码基因的共表达信息[36];RiceATM是用来挖掘miRNA与水稻农艺性状关系的数据库,包括187个水稻品种的8个产量相关性状数据集和193个miRNAs在这些品种中的表达水平和靶基因数据集[37]。一些植物综合ncRNA数据库,也包含比较丰富的水稻ncRNA信息。CSRDB是较早开发的sRNA数据库,整合了水稻和玉米的sRNA和它们的靶基因信息[38];miRbase是广泛使用的miRNA数据库,包括已知的水稻miRNA的成熟序列、前体序列和靶基因信息[39];PceRBase整合了包含水稻在内的26个植物物种的竞争性内源RNA(Competing endogenous RNA,ceRNA)数据集,为miRNA−靶基因调控网络的研究起到了补充作用[40]。此外,GreeNC[41]、PLncDB[42]、CANTATAdb[42-43]、PmiREN[44]和PlantcircBase[45]等多物种数据库中也包含水稻lncRNA、circRNA和miRNA信息。
表 2 水稻转录和转录后调控相关数据库Table 2. The transcriptional and posttranscriptional regulation related databases in rice数据库
Database描述
Description参考文献
ReferenceRiceXPro 微阵列数据集,自然条件下各个生长发育阶段、幼苗激素和胁迫处理的基因表达信息,https://ricexpro.dna.affrc.go.jp/ [27] CREP ‘珍汕97’和‘明恢63’的39个组织的基因表达信息,
http://crep.ncpgr.cn/crep-cgi/home.pl[28] RED 水稻9个组织,在不同生长阶段和处理的基因表达谱和基因共表达网络,http://expression.ic4r.org [29] TENOR 包括‘日本晴’在不同环境胁迫和激素处理条件下的140个mRNA-seq数据集,https://tenor.dna.affrc.go.jp/ [30] PPRD 11726个水稻mRNA-seq数据集,使用统一流程和最新的参考基因组进行分析和整合,http://ipf.sustech.edu.cn/pub/ricerna/ [31] eRice ‘日本晴’和‘9311’的mRNA-seq、DNA甲基化和组蛋白修饰数据库,http://www.elabcaas.cn/rice/index.html [32] RiceENCODE 综合调控RNA转录的DNA修饰、组蛋白修饰、染色质构象等表观调控元件,http://glab.hzau.edu.cn/RiceENCODE/ [33] RiceNCexp 提供基于mRNA-seq和sRNA-seq的基因和sRNA转录水平和共表达网络信息,https://cbi.njau.edu.cn/RiceNCexp/ [34] ARMOUR 7个水稻品种在不同发育时期、组织和胁迫下的miRNA和相应的靶标信息,https://www.icgeb.org/armour.html [35] RiceLncPedia 包含了水稻ncRNAs的表达谱、变异位点、ncRNA之间和ncRNA与编码基因的共表达网络信息,http://3dgenome.hzau.edu.cn/RiceLncPedia [36] RiceATM 挖掘miRNA与水稻农艺性状的关系,包括表型选择、样本分组、微阵列数据预处理、统计分析和靶基因预测等功能,http://syslab3.nchu.edu.tw/rice/ [37] CSRDB 整合了水稻和玉米的sRNA和它们的靶基因信息,http://sundarlab.ucdavis.edu/smrnas/ [38] miRbase 包含水稻已知和新的miRNA的序列和前体序列信息,是使用最广泛的miRNA综合数据库,http://mirbase.org/ [39] PceRBase 包含水稻等26个物种的ceRNA、miRNA和它们的靶基因信息,http://bis.zju.edu.cn/pcernadb/index.jsp [40] GreeNC 2.0 水稻lncRNA数据库,http://greenc.sequentiabiotech.com/wiki2/Main_Page [41] PLncDB 提供lncRNA的长度、类型、表达谱和表观遗传等信息,http://plncdb.tobaccodb.org/ [42] CANTATAdb 提供lncRNA长度、类型和表达谱信息,http://cantata.amu.edu.pl,http://yeti.amu.edu.pl/CANTATA/ [43] PmiREN 包含水稻miRNA及其前体序列、二级结构、表达模式、潜在靶点等信息,http://www.pmiren.com/ [44] PlantcircBase 提供水稻circRNA的分类、表达谱信息,http://ibi.zju.edu.cn/plantcircbase/ [45] PlaASDB 水稻和拟南芥在非生物和生物胁迫下的AS事件及AS与基因表达之间的联系,http://zzdlab.com/PlaASDB/ASDB/index.html [46] PlantAPAdb 水稻和拟南芥等6个物种基因组范围内的APA位点及注释信息,http://www.bmibig.cn/plantAPAdb [47] 选择性多聚腺苷酸化 (Alternative cleavage and polyadenylation, APA)是指具有多个多聚腺苷酸化信号位点(Polyadenylation signal, PAS) 的基因在mRNA 加工过程中,由于可变剪切(Alternative splicing,AS) 和APA调控过程,形成不同的mRNA异构体的现象[48]。PlaASDB综合分析了水稻和拟南芥在生物和非生物胁迫条件下的AS事件,并将AS与差异表达基因进行了关联[46]。PlantAPAdb基于3′-seq(3′ sequencing),在基因组范围内详细鉴定了水稻和拟南芥等6个物种的APA位点,并提供了多样化的注释信息,包括APA在基因组位置、异质性切割位点、表达水平和样本信息等[47]。综上所述,转录和转录后调控过程中涉及的数据库,主要集中在基因表达谱和ncRNA方面,有关AS、APA和RNA修饰等转录后调控的数据库资源仍然较少。因此,亟需构建水稻RNA修饰等转录后调控数据库,为挖掘水稻优异性状和转录后调控之间的潜在联系提供数据基础。
1.3 基因网络数据库
在植物中,生物网络广泛用于功能基因的挖掘、基因功能和重要农艺性状遗传机制解析的研究中[49]。生物网络是以生物分子作为节点,分子之间的相互作用作为连接边构成的系统,比较常见的有蛋白质互作网络、基因调控网络、共表达网络、代谢网络等[50]。在植物中,基因网络推断主要依赖于表达数据,使用相关性分析和互信息算法推断2个基因之间的潜在关系[51]。近年来,利用机器学习算法,如图卷积神经网络 (Graph convolutional network, GCN),能够有效降低网络的冗余,得到节点数尽可能少,并能最大化地表示原始图的子图,从而更真实地反映基因之间的互作网络[52]。目前已开发的水稻基因网络数据库主要是基于转录组和代谢组数据集构建的,涉及的网络推断方法主要有相关性分析、互信息算法、同源映射和机器学习(表3)。RiceFREND和OryzaExpresss是较早建立的基因网络数据库,二者均是基于微阵列数据集构建的基因共表达网络,并提供了比较友好的网络交互界面和灵活的数据检索和查询功能[53-54];RiceAntherNet提供了水稻花粉和花药发育过程的基因网络信息,包括57个水稻花药组织微阵列数据集,有助于挖掘花药和花粉调控网络中的关键基因[55];NetREx是基于已发表的mRNA-seq数据集构建的不同胁迫条件下水稻基因共表达网络数据库,可以实现胁迫或组织特异性共表达模型的查看,并关联了基因功能注释和基因通路等数据集[56]。此外,NetREx还提供了水稻、拟南芥、小麦、玉米、大麦和高粱的直系同源基因信息;PRIN是基于同源映射法,通过对果蝇、线虫和拟南芥等模式物种蛋白互作数据进行同源推断获得的第1个水稻蛋白质互作网络数据库[57];RiceNet和RicePPINet是利用机器学习构建的基因网络数据库[58-59]。其中,RiceNet基于多组学数据集,利用贝叶斯框架、网络直接邻域法和机器学习整合微阵列、转录组、蛋白互作和不同物种的同源映射等数据集,构建了一个由25765个基因结点和1775000个连接边组成的基因网络[58]。RicePPINet包含了16895个蛋白质的708819个基于机器学习算法预测的蛋白质−蛋白质相互作用网络,并通过与已知蛋白互作信息和试验证明了预测结果的可靠性[59]。
表 3 已发表的水稻基因网络数据库Table 3. The publised gene network databases in rice数据库
Database描述
Description参考文献
ReferenceRiceFREND 基于不同组织不同生长发育阶段的微阵列数据构建的共表达网络,http://ricefrend.dna.affrc.go.jp/ [53] OryzaExpress 基于微阵列数据构建的共表达网络,http://plantomics.mind.meiji.ac.jp/OryzaExpress/ [54] RiceAntherNet 基于微阵列数据集构建的花粉和花药发育过程的共表达网络,https://www.cpib.ac.uk/anther/riceindex.html [55] NetREx 基于同源映射和转录组数据集构建的基因在逆境和激素处理下的共表达网络,https://bioinf.iiit.ac.in/netrex/index.html [56] PRIN 基于模式物种蛋白互作基因同源映射构建的水稻蛋白质互作网络,http://bis.zju.edu.cn/prin/ [57] RiceNetv2 基于蛋白互作、mRNA-seq等多种数据集,利用机器学习算法构建的基因网络,http://www.inetbio.org/ricenet [58] RicePPINet 基于机器学习算法构建的蛋白质互作网络,http://netbio.sjtu.edu.cn/riceppinet [59] 在植物中,基因网络推断主要依赖于表达数据,使用相关性分析和互信息算法推断两个基因之间的潜在调控关系。严格来说,基于表达数据推断出的基因与基因之间的联系并不一定是客观存在的。机器学习算法可以用来整合不同类型的基因网络数据集,如基因共表达网络、蛋白互作网络和代谢网络,尽可能准确地推断基因之间的关系。此外,利用GCN算法可以降低网络的冗余,得到节点数尽可能少并能最大化表示原始图的子图。综上所述,基于多组学数据和机器学习算法构建的基因网络,将更加准确地反映基因之间的相互作用关系。
1.4 种质资源信息数据库
随着大规模种质资源测序项目的开展,积累了海量的数据集。研究者通过构建种质资源信息数据库整合和共享这些数据集,并为数据的进一步挖掘和利用提供了在线分析工具。在水稻中,已有多个种质资源信息数据库,如MBKBASE、RiceVarMap和SNP-Seek,它们为水稻分子育种提供了丰富的基因组变异、单倍型以及变异的遗传力等信息(表4)。MBKBASE收录了130578份水稻种质资源信息,并以‘R498’和‘日本晴’为参考基因组,进行了单倍型鉴定,并提供了单倍型与性状和基因表达水平的关联信息[60];RiceVarMap收录了全球140个国家和地区的4726份水稻种质资源及其遗传变异信息,包括14541446个SNP和2855580个InDel[61];SNP-Seek包含了来源于119个国家的4036份水稻品种资源信息,包括种质资源群体结构、遗传变异和57个表型等数据集[62];SR4R提供了5152份水稻资源的变异图谱,通过统一的流程整合遗传变异数据,有效地减少了SNP冗余[63];HapRice包括76份世界水稻品种的3334个SNP和177份日本水稻品种的3252个SNP,并构建了SNP单倍型图谱[64];RFGB基于3KRG数据集,整合了表型、单倍型、SNP和InDel等信息[65];中国水稻品种及其系谱数据库是国家水稻数据中心(http://www.ricedata.cn/) 的一个子平台,主要收录省级以上审定品种、大面积推广品种、外引品种以及地方性农家品种,具有品种检索和系谱追溯2个核心功能[66]。转座子是重复序列的一种,也是一种基因组变异,在真核生物中广泛存在。水稻种质资源转座子注释数据库有RiTE DB[67]和RTRIP[68]。RiTE DB是第1个水稻转座子数据库,包括从266份水稻品种中鉴定到的54911个转座子信息[67];RTRIP包含基于3KRG的基因组重测序数据构建的转座子序列位点图谱[68]。
表 4 已发表的水稻种质资源信息数据库Table 4. The published germplasm information resources in rice数据库
Database描述
Description参考文献
ReferenceMBKBASE 以‘R498’和‘日本晴’为参考基因组,包含130578份种质的表型、群体结构和单倍型信息,https://www.mbkbase.org/rice/ [60] RiceVarMap 4726份水稻种质资源的基因组变异、基因型和表型数据,http://ricevarmap.ncpgr.cn/v2/ [61] SNP-Seek 4036份水稻的SNP图谱、表型和全基因组关联分析数据集,https://snp-seek.irri.org/ [62] SR4R 包含5152份种质资源的遗传变异、群体遗传学和进化基因组学数据集,http://sr4r.ic4r.org/ [63] HapRice 253份国际和日本来源的水稻种质的SNP和单倍型图谱,http://qtaro.abr.affrc.go.jp/index.html [64] RFGB 3KRG的表型、遗传变异和基因单倍型信息,http://www.rmbreeding.cn/ [65] RiceData 包含省级以上审定品种、大面积推广品种、外引品种以及地方性农家品种信息,http://www.ricedata.cn/variety/ [66] RiTE DB 包含266份水稻品种中鉴定到的54911个转座子信息数据集,https://www.genome.arizona.edu/cgi-bin/rite/index.cgi [67] RTRIP 3KRG的基因组转座子信息,提供转座子序列位点图谱、遗传多样性、基因组进化和分子标记等信息,http://ibi.zju.edu.cn/Rtrip/index.html [68] 在作物驯化和遗传改良过程中,遗传多样性不断变窄。国家/国际基因库(种质库)提供了丰富来源的等位基因,这些等位基因可能对作物遗传改良至关重要。因此,在基因组和表型组水平上对尽可能多的种质资源进行鉴定,为确定等位基因与表型的联系提供了信息,从而有助于育种决策。综上所述,对基因库中存在的全部种质进行测序和表型鉴定,获得所有种质的遗传变异、等位基因以及它们与表型的关联信息,构建水稻种质资源综合数据库是未来育种计划的重要组成部分。
2. 水稻基因编辑生物信息工具
2.1 基因编辑系统概述
基因编辑能在不引入DNA双链断裂 (Double-strand DNA breaks, DSBs)的情况下,对靶位点一定范围内的碱基进行编辑。CRISPR/Cas系统作为第3代基因组编辑技术,具有操作简单灵活、精准靶向和效率高等优点,在基础研究和生物育种中具有广泛的应用前景[69-70]。
如表5所示,主要的基因编辑系统包括胞嘧啶碱基编辑器(Cytosine base editor, CBE)、腺嘌呤碱基编辑器(Adenine base editor, ABE)、胞嘧啶−鸟嘌呤碱基编辑器 (Cytosine-to-guanine base editor, CGBE)、引导编辑器(Prime editor,PE)等,已广泛应用在人类、动植物和微生物的相关研究中[71]。近年来,研究者将Cas9蛋白和反转录酶蛋白 (M-MLV)融合,以及将sgRNA改造成可提供修复模板的pegRNA,构建了PE系统,实现了A、T、C、G碱基的自由替换和小DNA片段的替换或插入,可编辑远离PAM(Protospacer adjacent motif)的序列,显著提高了基因编辑的精准度[72]。
表 5 常用的基因编辑系统Table 5. Commonly used gene editing systems编辑系统
Editing system编辑作用
Editing function用途
ApplicationCRISPR/Cas 靶点切割/突变 基因敲除、遗传改良 CRISPR/Cas/Donor DNA片段插入/替换 DNA片段突变、遗传改良 CRISPR/Cas-CBE 单碱基转换:C > T(G > A) 单碱基转换、遗传改良 CRISPR/Cas-ABE 单碱基转换:A > G(T > C) 单碱基转换、遗传改良 CRISPR/Cas-CGBE 单碱基转换:C > G(G > C) 单碱基转换、遗传改良 CRISPR/Prime editors 小片段插入、替换 单碱基的任意转换和小DNA片段的
替换或插入、遗传改良2.2 基因编辑靶标设计生物信息工具
基因编辑中目标基因靶位点的选择、sgRNA(Single guide RNA)表达盒的设计和组装、靶位点片段的扩增与测序、脱靶(Off-target)及自靶向(Self-target)分析等过程仍会耗费大量的人力物力。近年来,利用生物信息工具辅助基因编辑的设计和分析流程,可在减少人为错误的同时也极大地节省时间和人力成本,使基因编辑高通量化[73-75]。
表6列出了已开发的水稻基因编辑数据库或在线工具。CRISPR-GE提供了“一站式”基因组编辑便捷工具包,包含了靶点筛选、引物设计、结果鉴定、脱靶分析等流程的在线分析软件[73];CRISPR-GE持续更新,在后续的开发中添加了利用微同源末端连接介导基因组大片段删除的靶点设计工具MMEJ-KO、辅助碱基编辑靶点选择工具BEtarget、基于多个高质量水稻基因组整合的数据库GeneCat、以及基于二代测序方法高通量分析靶位点编辑情况的HiDecode[73-74,76-78];此外,CRISPR REGN提供基因编辑在线工具包,如sgRNA设计工具Cas-Designers[79]、脱靶预测工具Cas-Offinder[80]和预测MMEJ(Microhomology-mediated end joining)介导片段删除的效率分析工具Microhomology-Predictor等[75];CRISPR-P支持49个物种基因组的sgRNA设计,开发了靶向效率和脱靶概率评估系统,使预测结果更为准确,并支持多种CRISPR/Cas系统的设计[81-82];CRISPRbase收集了包括水稻在内的17个物种多达120万条碱基编辑信息,整理分析了多个编辑系统在不同物种中的编辑效率、靶点偏好性和精准度,并提供了在线分析工具[83];PGED收集了水稻等8个物种的CRISPR/Cas9编辑植株信息,包括sgRNA序列、序列变异和表型等信息,用户可以通过物种或基因名查找已有的CRISPR/Cas9突变体[84]。
表 6 水稻基因编辑生物信息工具与数据库Table 6. The bioinformatics tools and databases for gene editing in rice数据库
Database描述
Description参考文献
ReferenceCRISPR-GE “一站式”基因编辑设计工具包,涵盖靶点筛选、引物设计、编辑结果鉴定、脱靶分析等流程,http://skl.scau.edu.cn/ [73-74] MMEJ预测工具 评估MMEJ介导片段删除效率,http://www.rgenome.net/ [75] BEtarget 提供候选靶点在基因上的位置、GC含量、潜在脱靶值和脱靶位点、编辑窗口内的碱基变化及对应的氨基酸变化等信息,http://skl.scau.edu.cn/betarget/ [76] MMEJ-KO 基于微同源删除基因组片段的靶点设计工具,http://skl.scau.edu.cn/mmejko/ [77] GeneCat 快速提取基因组序列工具,http://skl.scau.edu.cn/genecat/ [78] Cas-Designers sgRNA设计线上工具,http://www.rgenome.net/cas-designer/ [79] Cas-Offinder sgRNA脱靶预测线上工具,http://www.rgenome.net/cas-offinder/ [80] CRISPR-P 多功能的sgRNA设计工具,随后升级为CRISPR-P 2.0,支持49个物种基因组的sgRNA设计,http://crispr.hzau.edu.cn/CRISPR2/ [81-82] CRISPRbase 碱基编辑综合知识平台,统计了多个物种的编辑效率、靶点偏好性和精准度,并进行功能注释,http://crisprbase.maolab.org/ [83] PGED 植物基因组编辑数据库,可供用户查阅或上存基因编辑靶位点、突变情况、表型信息等,http://plantcrispr.org [84] CAFRI-Rice 水稻冗余基因数据库,为基因编辑候选靶标的选择提供参考,http://pcafri-rice.khu.ac.kr [85] Ribo-uORF uORF综合数据库,收集了6个物种的高可信度uORF和TIS位点信息,为uORF编辑提供靶标,http://rnainformatics.org.cn/RiboUORF [86] CAFRI-Rice基于系统发育分析和基因表达模式,建立了水稻冗余基因数据库,为基因编辑候选靶标的选择提供了参考[85]。uORF (Upstream open reading frame, uORF)定义为位于5′UTR上游的开放读码框,对主开放读码框(Primary ORF, pORF)的翻译具有抑制作用[86]。通过对uORF区域进行基因编辑,能够实现对目标蛋白表达量的精准调控[87]。Liu等[86]基于深度测序Ribo-seq(Ribosome profiling)和QTI-seq(Quantitative translation initiation sequencing),构建了Ribo-uORF数据平台,包含了6个物种的501554条高可信度的uORF和107914条翻译起始位点 (Translation initiation site, TIS)信息。Ribo-uORF提供了丰富的uORF信息,为通过基因编辑精准调控蛋白质的表达水平提供了候选靶标[86]。
综上所述,生物信息工具辅助基因编辑可以有效提高编辑精准度(降低潜在脱靶风险)、预测编辑效果和提供新型编辑靶点。然而,全基因组测序结果发现了编辑植株中存在许多与序列相似度无关的脱靶位点。开发生物信息工具,对与序列相似度无关的脱靶事件进行预测,有助于进一步提高编辑精准度。此外,结合uORF、dORF(Downstream ORF)和m6A修饰等数据库,将有助于指导新型编辑策略的开发与应用。
3. 水稻智能育种生物信息工具
随着人类社会步入互联网、大数据和人工智能“三位一体”时代,对育种提出了革命性理念,即强调生命科学、信息科学与育种科学的深度融合[2-3,88]。在智能育种阶段,育种专家将综合多层面生物技术与信息技术推动育种向着智能化的方向快速发展:1)利用高通量低成本的基因组测序技术和表型鉴定技术,结合人工智能图像识别技术,通过基因型与表型数据的自动化获取与解析,实现组学大数据的快速积累;2)利用生物信息学与机器学习的方法和手段,整合遗传变异和各类组学数据、杂交育种数据,实现控制作物性状关键调控基因的快速挖掘与表型的精准预测;3)利用基因编辑与合成生物学技术,通过人工改造基因元器件与人工合成基因网络,实现作物具备新的抗逆、高效和优良生物学性状;4)利用作物组学大数据与人工智能技术,建立机器学习预测模型,在全基因组层面上建立智能组合优良等位基因的自然变异、人工变异、数量性状位点的育种设计方案,实现智能、高效、定向培育新品种。
由于海量的植物育种相关的大数据必须借助于人工智能进行数据分析、处理和预测,机器学习模型在智能育种中起重要作用。表7列出了基于机器学习进行组学数据处理和辅助育种决策的软件和算法,可分为线性和非线性的方法。线性模型以基于混合线性模型的方法为代表,如基因组最佳线性无偏预测(Genomic best linear unbiased prediction,GBLUP)、岭回归最佳线性无偏预测 (Ridge regression best linear unbiased prediction, rrBLUP)、贝叶斯算法(BayesA/B)等;非线性模型主要以随机森林(Random forest, RF)、支持向量机(Support vector machine, SVM)和神经网络(Neural network)为主。目前存在3类基于不同的统计模型的智能育种工具:第1类是免费开源的包/库,如rrBLUP和sommer等R包,以及基于Python的机器学习模块,如sklearn模块中的RF和SVM算法;第2类是成熟的遗传评估软件,主要用于动物育种,但也有少量功能适用于植物育种,如JWAS;第3类是基于网页或图形界面工具,例如solGS和IPAT等[106]。在智能育种设计方案方面,Xu等[107]提出了一种名为iGEP的智能化育种方案,综合运用多组学信息、大数据技术和人工智能算法,通过利用从多个来源获取的数据(包括时空组学,基因组学、表型组学和环境组学),建立智能育种机器学习模型,并通过合成生物学的方法从头培育新品种,实现理想作物的智能化育种。
表 7 可用于水稻智能育种的机器学习软件和算法Table 7. The machine learning software and algorithms for intelligent breeding in rice软件
Software模型
Model描述
Description参考文献
ReferenceBGLR BL、BR、BayesA/B 基于基因−环境互作和多性状的基因组选择模型的构建,https://github.com/gdlc/BGLR-R [89] BRNN brnn 基于双向循环神经网络进行基因组选择和表型预测,https://cran.r-project.org/web/packages/brnn/ [90] BWGS BayesA/B、BL、BRR 基于R语言开发进行基因组选择和表型预测,https://cran.r-project.org/web/packages/BWGS/ [91] CropGBM LightGBM 基因型和表型数据预处理、群体结构分析、SNP 特征选择、表型预测和数据可视化,https://github.com/YuetongXU/CropGBM [92] DeepGS CNN 基于深度学习整合多组学数据进行基因组选择,https://github.com/cma2015/DeepGS/ [93] DNNGP DNN 基于深度神经网络整合多组学数据进行基因组选择,http://github.com/AIBreeding/DNNGP/ [94] HIBLUP BLUP 利用谱系、基因组和表型信息,评估个体的遗传价值,https://hiblup.github.io/ [95] KAML KAML、GBLUP 控制质量性状和数量性状的关键基因挖掘,https://github.com/YinLiLin/KAML [96] PopVar RRBLUP、 BayesA/B/C、
BL、BRR利用基因型和表型数据预测双亲后代的遗传方差和表型值,https://cran.r-project.org/web/packages/PopVar/ [97] rrBLUP RRBLUP 基因组分子标记遗传效应估计与表型预测,https://cran.r-project.org/web/packages/rrBLUP/ [98] sommer GBLUP、RRBLUP 加性效应、显性效应、上位性效应评估和遗传力的计算,https://cran.r-project.org/web/packages/sommer/ [99] STGS ANN、BLUP、LASSO、
RF、RR、SVM基于分子标记对单一性状进行基因组选择,https://cran.r-project.org/web/packages/STGS/ [100] GCTA BLUP SNP 遗传力评估和全基因组关联分析,http://cnsgenomics.com/software/gcta/ [101] JWAS Bayes 基因组选择和全基因组关联分析,http://reworkhow.github.io/JWAS.jl/latest/ [102] PIBULP BLUP 遗传参数评估与育种值估计,https://github.com/huiminkang/PIBLUP [103] solGS RRBLUP 基于组学数据进行复杂性状表型预测,http://cassavabase.org/solgs [104] IPAT GBLUP、RRBLUP、BayesB 全基因组关联分析与育种值估计的在线图形化界面工具,http://poissonfish.github.io/iPat/index.html [105] 智能育种技术具有广阔的应用前景,是未来水稻育种的发展方向之一。目前智能育种发展仍处于起步阶段,面临多组学大数据积累不足、基因分型成本较高、基因组预测育种模型原创性不足、种质资源较为分散等众多挑战[108]。因此,制定多维度数据采集、分析、存储与管理的标准与规范,协同建立通用的育种大数据平台,实现育种信息的充分共享与利用,是现阶段促进生物信息学在智能育种中应用的首要目标。
4. 总结与展望
自从第1个水稻‘日本晴’参考基因组公布以来,生物信息学在组学数据分析与整合、基因挖掘、基因网络推断和重要性状遗传解析等方面起到了关键性作用。基于海量的数据集,研究者们开发了许多种类丰富的生物信息数据库和在线工具。由于数据来源、试验设计和分析方法的不同,不同数据库资源很难整合。因此,需要系统整合不同类型数据库,开发大型水稻综合数据库平台,以供不具有高级生物信息学技能的生物学家有效使用。此外,如何将多组学数据与表型数据进行整合,指导基因组选择和全基因组设计育种,提高水稻产量、品质和环境适应性,是从事相关科研工作的研究者需要考虑的问题之一。在数据库类型方面,涉及基因转录后调控、基因编辑和智能育种的资源仍然缺乏。此外,修饰组和翻译组,如RNA甲基化、蛋白质泛素化和uORF等可以实现基因表达水平的精准调控,开发相应的数据库和分析工具,将有利于分子设计育种和品种的定向改良。最后,鉴于大部分数据库更新周期较长或没有更新,应用机器学习的方法,如文本挖掘、随机森林和图卷积神经网络,对最新发布的数据集进行自动下载,并整合到后台数据库,将有望实现数据库的实时动态更新。
-
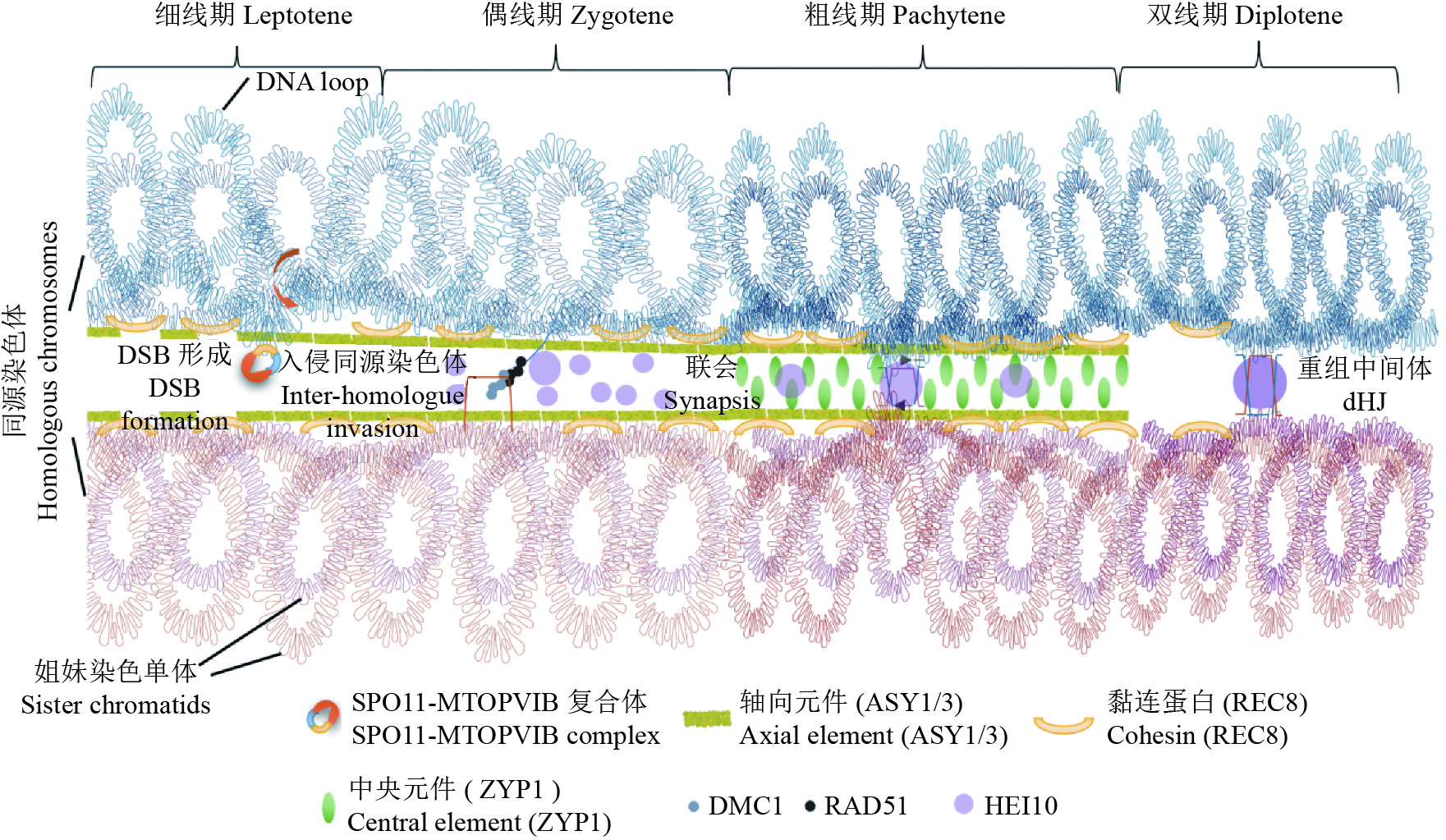
图 1 植物减数分裂染色体的牵回环–轴模型
减数分裂同源染色体的配对和联会是重组的重要保证。在减数分裂细线期,联会起始于轴向元件ASY1和ASY3在染色体上的加载,和黏连蛋白REC8一起最终形成一个线性的轴结构,使得姐妹染色单体沿着轴形成一个个DNA环。在细线期和偶线期转换时,DNA环会被牵拉到染色体轴上,由SPO11-MTOPVIB复合体介导双链断裂(DSBs)的产生。随后在偶线期,3′单链DNA会在RAD51-DMC1的介导下入侵到同源染色体间,形成D-loop,从而促进同源染色体配对。在粗线期,中央元件ZYP1加载到一对同源染色体轴中间,最终形成联会复合体的完整结构。在此时,大部分的DSB被修复,少量形成了重组中间体的结构。最后在双线期,联会复合体开始分解,同源染色体分离,只留下重组的区域形成交叉(Crossovers)
Figure 1. The model of tethered loop-axis of meiotic chromosomes in plant
The pairing and synapsis of meiotic homologous chromosomes are important for recombination. At leptotene, the synapsis initiates with the loading of axial elements ASY1 and ASY3 on the chromosome, which eventually forms a linear axis structure together with cohesin REC8. Sister chromosomes form DNA loops to array along the axis. During leptotene-diplotene transition, DNA loops are tethered onto the chromosome axis, and double-strand breaks (DSBs) are induced by SPO11-MTOPVIB complex. Subsequently, 3′ single-strand DNA end searches the homologous chromosome by RAD51-DMC1 to form a D-loop, thus promoting homologous chromosome pairing. At pachytene, the central element ZYP1 is loaded onto the centre of a pair of homologous chromosome axes, and finally forms the complete synaptonemal complex (SC) with axial elements. At this point, most DSBs are repaired, and a small amount forms recombination intermediates. At diplotene , following SC disassembly, homologs are separated except where crossovers have formed
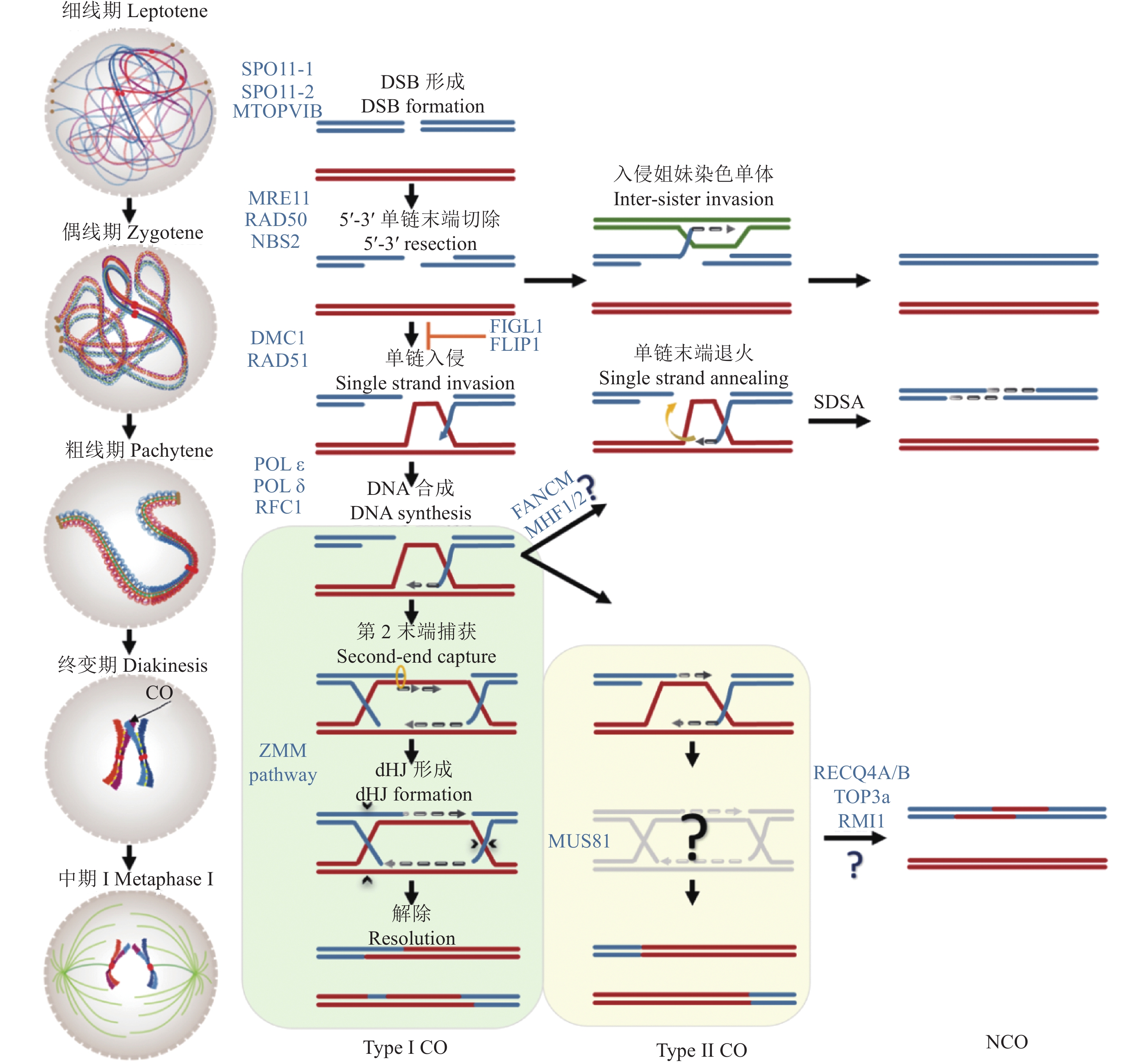
图 2 植物减数分裂重组修复途径模型
减数分裂的重组起始于双链断裂的形成,由SPO11-MTOPVIB复合体介导。双链断裂末端随后被切割处理形成3′单链尾巴。双链断裂修复可以选择以姐妹染色单体为模板,也可以在RAD51-DMC1的帮助下使3′单链末端入侵同源染色体形成置换环结构进行修复,后者形成减数分裂染色体重组。在以同源染色体为模板的修复中,DNA合成,第2链末端捕获和链接最终形成重组中间体的经典结构——双Holliday交叉,并最终解除形成干涉敏感型交换。同时,还存在着重组蛋白MUS81依赖的干涉不敏感型交换,但是目前植物中的重组中间体以及产物不太清楚。此外,在单链入侵后,当第2末端无法捕获时还存在一条合成依赖的链退火途径,最终也以姐妹染色单体为模板进行修复,并会产生非交换。在重组通路中,还存在着3条不同的重组抑制通路:FIGL1-FLIP、FANCM-MHF1/2和RECQ4A/B-TOP3α-RMI1,都参与抑制MUS81依赖的干涉不敏感型交换途径,并促进合成依赖的链退火
Figure 2. The model for meiotic recombination in plant
Meiotic recombination initiates with the formation of DSBs, which are mediated by the SPO11-MTOPVIB complex. DSB ends are cleaved to form 3′ single-strand tails. Subsequently, DSB can be repaired by selecting the sister chromatids as the template. Alternatively, 3′ single-strand ends invade the homolog to form a D-loop for repair, which is known as recombination. Following the repair progress, DNA synthesis, second strand end capture and ligation lead to the formation of double Holliday junctions (dHJs), which are the classical structure of the recombination intermediates and finally resolved as ZMM-dependent interference-sensitive crossovers (Type I COs). Meanwhile, MUS81-dependent pathway results in interference-insensitive crossovers (Type II CO) , but the recombination intermediates and their products in plants are not well understood. In addition, single-strand invasion can be processed by synthesis-dependent strand annealing pathway (SDSA), and chooses the sister chromatids as the template for repair to produce NCO. During meiotic recombination, there are also three different recombination inhibitory pathways, including FIGL1-FLIP, FANCM-MHF1/2 and RECQ4A/ B-Top3α-RMI1, which are involved in the inhibition of MUS81-dependent Type II CO and promote SDSA
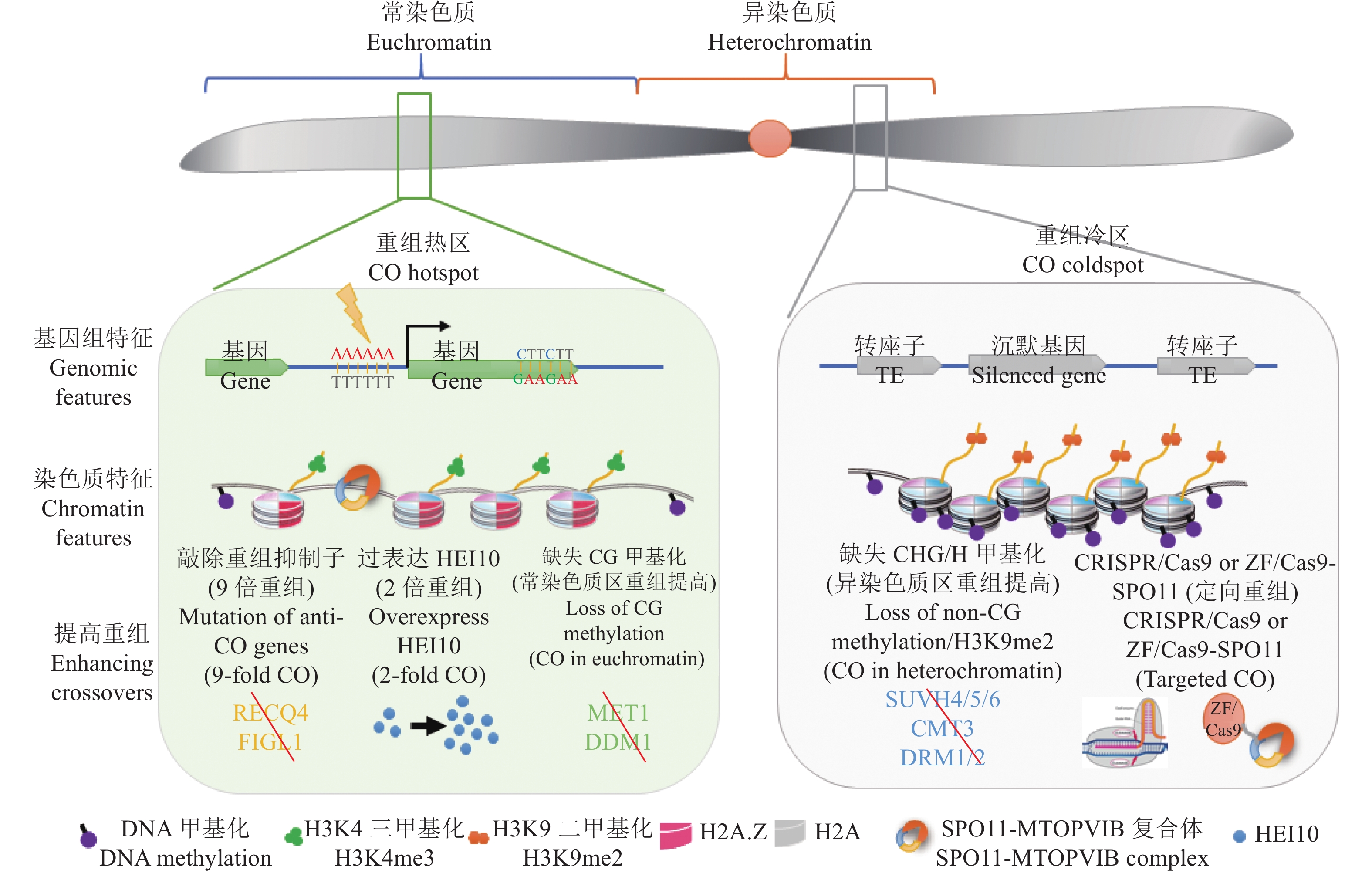
图 3 拟南芥减数分裂重组(CO)热点的基因组/染色质特征和调控模型
减数分裂重组在染色体上不是均匀分布的。通常倾向于发生在常染色质区,并且DSB通常发生于基因的转录起始位点或转录终止位点。重组热区伴随着低核小体密度、H2A.Z和H3K4me3的富集,并且含有AT-rich 和CTT 基序。而在异染色质区通常结构致密,TE和重复序列富集,伴随高非CG甲基化和H3K9me2,这些都是重组的抑制因素。在常染色质区提高减数分裂重组的方法有:突变重组抑制子、过表达重组酶HEI10和突变MET1和DDM1;而在异染色质区提高重组通常通过降低非CG甲基化或H3K9me2,另外还可以通过定向重组的方法提高目标区域重组
Figure 3. The model for regulation of Arabidopsis meiotic crossover hotspots by genomic and chromatin features
The distribution of meiotic recombination events is not uniformly along chromosomes. The crossovers (COs) generally tends to occur in euchromatin regions, and DSBs usually occur in transcription start sites (TSS) or transcription stop sites (TTS). Meiotic recombination hotspots display low nucleosome density, occupancy of H2A.Z and H3K4me3 enrichment with AT-rich and CTT motifs. Heterochromatin is highly compacted with lots of TE and repeat sequences accompanied by high non-CG methylation and H3K9me2, which are the inhibitors of recombination. The approaches for improving meiotic recombination frequency on euchromatin are as follows: Mutating anti-CO genes, overexpressing the recombinase HEI10 or mutating DNA methyltransferases MET1 and remodeler DDM1. Increasing COs on heterochromatin can be achieved by reducing non-CG methylation or H3K9me2, and targeted recombination
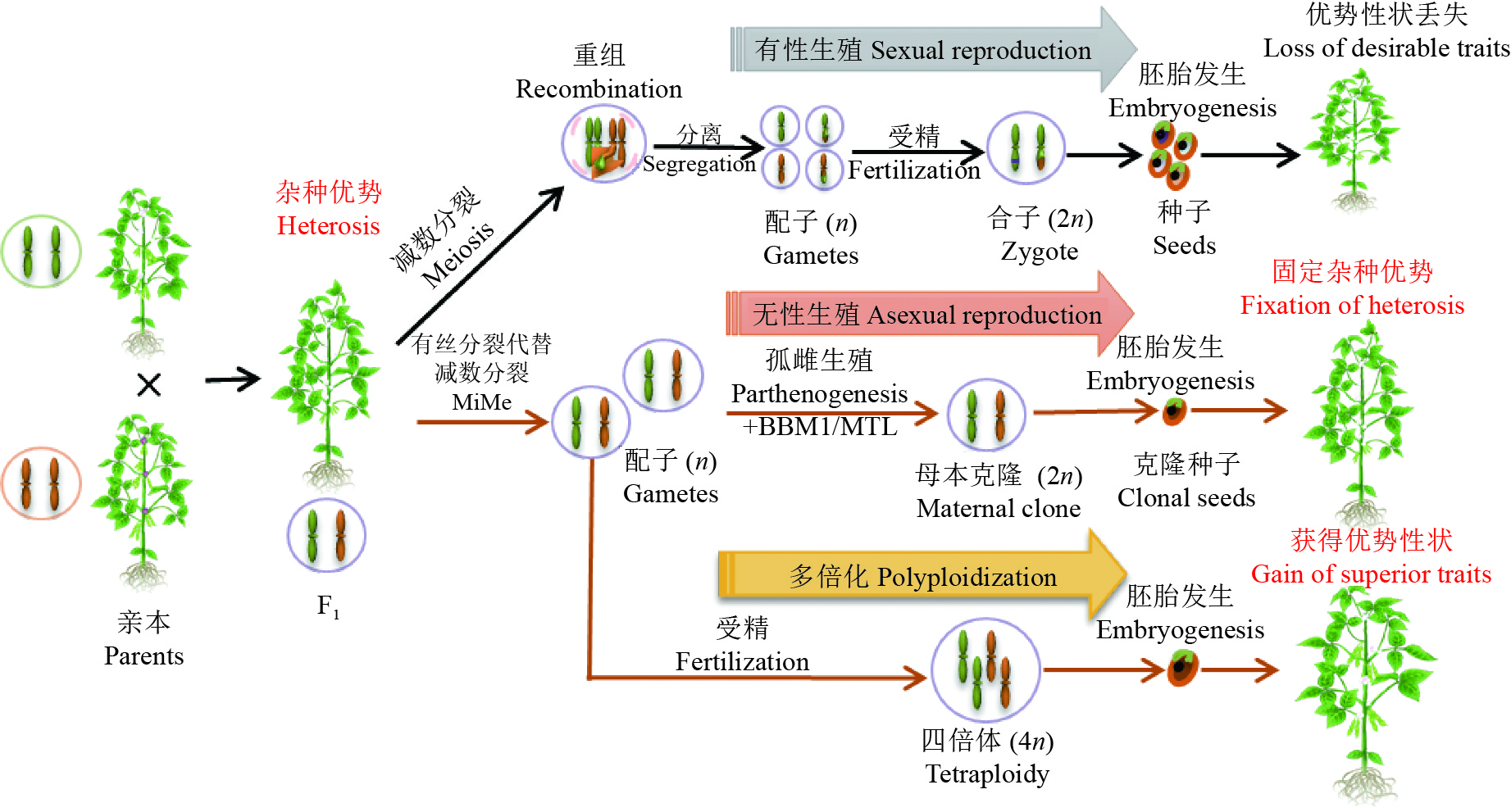
图 4 操纵减数分裂创新作物种质
植物杂交后代具有较亲本优越的表型或适应性称为杂种优势。在有性生殖过程中,F1代杂交植物经历减数分裂染色体重组和分离,导致配子间的等位基因再分配。而后植物双受精使雄配子与雌配子融合,由于减数分裂重组,在F1代中观察到的理想杂种优势性状通常在其后代中丢失。与此相反,无融合生殖依赖于未减数孢子分裂、孤雌生殖和不依赖受精的功能性胚乳的形成。在改良的植物无融合生殖中,通过对有丝分裂代替减数分裂(Mitosis instead of Meiosis, MiMe)和BBM1/MTL基因的遗传操作,分别诱导有丝分裂代替细胞分裂,产生二倍体配子,而后通过孤雌生殖实现种子克隆。克隆种子与F1杂交植株基因一致,可以在后代中保持杂种优势。此外,还可以通过MiMe实现后代多倍化,从而提高后代植物重组频率,整合并增强优良性状
Figure 4. Manipulating meiosis for crop improvement
Heterosis is a phenomenon among progenies of plants, which is known as that hybrid plants can have superior traits compared to their parents. In the process of sexual reproduction, F1 hybrid plants undergo meiotic recombination and chromosome segregation, leading to redistribution of alleles among gametes. Then plant double fertilization is the fusion of male gametes and female gametes. Due to meiotic recombination, the desirable traits observed in F1 hybrid plants are usually lost in their progenies. In contrast, apomixis relies on the apomeiosis, parthenogenesis and autonomous endosperm. In the improved apomixis of plant, MiMe induced by genetic manipulation produces diploid gametes and misexpression of BBM1/MTL in egg cell triggers parthenogenesis, thereby producing the clonal reproduction through seeds. The genome of cloned seeds is consistent with F1 hybrid plants, which can maintain heterosis in the progeny. In addition, diploid gametes can be achieved through MiMe to produce polyploid offspring, which may improve the frequency of recombination and enhance superior traits
-
[1] BLAT Y, PROTACIO R U, HUNTER N, et al. Physical and functional interactions among basic chromosome organizational features govern early steps of meiotic chiasma formation[J]. Cell, 2002, 111(6): 791-802. doi: 10.1016/S0092-8674(02)01167-4
[2] ARMSTRONG S J, CARYL A P, JONES G H, et al. Asy1, a protein required for meiotic chromosome synapsis, localizes to axis-associated chromatin in Arabidopsis and Brassica [J]. Journal of Cell Science, 2002, 115(Pt 18): 3645-3655.
[3] FERDOUS M, HIGGINS J D, OSMAN K, et al. Inter-homolog crossing-over and synapsis in Arabidopsis meiosis are dependent on the chromosome axis protein AtASY3[J]. PLoS Genetics, 2012, 8(2): e1002507. doi: 10.1371/journal.pgen.1002507.
[4] KEENEY S, GIROUX C N, KLECKNER N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family[J]. Cell, 1997, 88(3): 375-384. doi: 10.1016/S0092-8674(00)81876-0
[5] VRIELYNCK N, CHAMBON A, VEZON D, et al. A DNA topoisomerase VI-like complex initiates meiotic recombination[J]. Science, 2016, 351(6276): 939-943. doi: 10.1126/science.aad5196
[6] SUGIMOTO-SHIRASU K, STACEY N J, CORSAR J, et al. DNA topoisomerase VI is essential for endoreduplication in Arabidopsis[J]. Current Biology, 2002, 12(20): 1782-1786. doi: 10.1016/S0960-9822(02)01198-3
[7] STACEY N J, KUROMORI T, AZUMI Y, et al. Arabidopsis SPO11-2 functions with SPO11-1 in meiotic recombination[J]. The Plant Journal, 2006, 48(2): 206-216. doi: 10.1111/j.1365-313X.2006.02867.x
[8] NEALE M J, PAN J, KEENEY S. Endonucleolytic processing of covalent protein-linked DNA double-strand breaks[J]. Nature, 2005, 436(7053): 1053-1057. doi: 10.1038/nature03872
[9] DE MUYT A, PEREIRA L, VEZON D, et al. A high throughput genetic screen identifies new early meiotic recombination functions in Arabidopsis thaliana[J]. PLoS Genetics, 2009, 5(9): e1000654. doi: 10.1371/journal.pgen.1000654.
[10] JI J H, TANG D, SHEN Y, et al. P31comet, a member of the synaptonemal complex, participates in meiotic DSB formation in rice[J]. Proceedings of the National Academy of Sciences of the United States of America, 2016, 113(38): 10577-10582. doi: 10.1073/pnas.1607334113
[11] WANG Y X, COPENHAVER G P. Meiotic recombination: Mixing it up in plants[J]. Annual Review of Plant Biology, 2018, 69: 577-609. doi: 10.1146/annurev-arplant-042817-040431
[12] MIAO C, TANG D, ZHANG H, et al. Central region component1, a novel synaptonemal complex component, is essential for meiotic recombination initiation in rice[J]. The Plant Cell, 2013, 25(8): 2998-3009. doi: 10.1105/tpc.113.113175
[13] PANIZZA S, MENDOZA M A, BERLINGER M, et al. Spo11-accessory proteins link double-strand break sites to the chromosome axis in early meiotic recombination[J]. Cell, 2011, 146(3): 372-383. doi: 10.1016/j.cell.2011.07.003
[14] LAMBING C, TOCK A J, TOPP S D, et al. Interacting genomic landscapes of REC8-cohesin, chromatin, and meiotic recombination in Arabidopsis[J]. The Plant Cell, 2020, 32(4): 1218-1239. doi: 10.1105/tpc.19.00866
[15] LAMBING C, KUO P C, TOCK A J, et al. ASY1 acts as a dosage-dependent antagonist of telomere-led recombination and mediates crossover interference in Arabidopsis[J]. Proceedings of the National Academy of Sciences of the United States of America, 2020, 117(24): 13647-13658. doi: 10.1073/pnas.1921055117
[16] LAM I, KEENEY S. Mechanism and regulation of meiotic recombination initiation[J]. Cold Spring Harbor Perspectives in Biology, 2014, 7(1): a016634. doi: 10.1101/cshperspect.a016634.
[17] UANSCHOU C, SIWIEC T, PEDROSA-HARAND A, et al. A novel plant gene essential for meiosis is related to the human CtIP and the yeast COM1/SAE2 gene[J]. The EMBO Journal, 2007, 26(24): 5061-5070. doi: 10.1038/sj.emboj.7601913
[18] OSMAN K, HIGGINS J D, SANCHEZ-MORAN E, et al. Pathways to meiotic recombination in Arabidopsis thaliana[J]. New Phytologist, 2011, 190(3): 523-544. doi: 10.1111/j.1469-8137.2011.03665.x
[19] WATERWORTH W M, ALTUN C, ARMSTRONG S J, et al. NBS1 is involved in DNA repair and plays a synergistic role with ATM in mediating meiotic homologous recombination in plants[J]. The Plant Journal, 2007, 52(1): 41-52. doi: 10.1111/j.1365-313X.2007.03220.x
[20] JI J H, TANG D, WANG M J, et al. MRE11 is required for homologous synapsis and DSB processing in rice meiosis[J]. Chromosoma, 2013, 122(5): 363-376. doi: 10.1007/s00412-013-0421-1
[21] JI J, TANG D, WANG K, et al. The role of OsCOM1 in homologous chromosome synapsis and recombination in rice meiosis [J] The Plant Journal, 2012, 72(1): 18-30.
[22] TSUBOUCHI H, OGAWA H. Exo1 roles for repair of DNA double-strand breaks and meiotic crossing over in Saccharomyces cerevisiae[J]. Molecular Biology of the Cell, 2000, 11(7): 2221-2233. doi: 10.1091/mbc.11.7.2221
[23] HU Q, TANG D, WANG H J, et al. The exonuclease homolog OsRAD1 promotes accurate meiotic double-strand break repair by suppressing nonhomologous end joining[J]. Plant Physiology, 2016, 172(2): 1105-1116.
[24] CHE L X, WANG K J, TANG D, et al. OsHUS1 facilitates accurate meiotic recombination in rice[J]. PLoS Genetics, 2014, 10(6): e1004405. doi: 10.1371/journal.pgen.1004405.
[25] WOLD M S. Replication protein A: A heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism[J]. Annual Review of Biochemistry, 1997, 66(1): 61-92. doi: 10.1146/annurev.biochem.66.1.61
[26] SHI B L, XUE J Y, YIN H, et al. Dual functions for the ssDNA-binding protein RPA in meiotic recombination[J]. PLoS Genetics, 2019, 15(2): e1007952. doi: 10.1371/journal.pgen.1007952.
[27] BROWN M S, BISHOP D K. DNA strand exchange and RecA homologs in meiosis[J]. Cold Spring Harbor Perspectives in Biology, 2014, 7(1): a016659. doi: 10.1101/cshperspect.a016659.
[28] AKLILU B B, SODERQUIST R S, CULLIGAN K M. Genetic analysis of the replication protein A large subunit family in Arabidopsis reveals unique and overlapping roles in DNA repair, meiosis and DNA replication[J]. Nucleic Acids Research, 2014, 42(5): 3104-3118. doi: 10.1093/nar/gkt1292
[29] ISHIBASHI T, KIMURA S, SAKAGUCHI K. A higher plant has three different types of RPA heterotrimeric complex[J]. The Journal of Biochemistry, 2006, 139(1): 99-104. doi: 10.1093/jb/mvj014
[30] MIAO Y J, SHI W Q, WANG H J, et al. Replication protein A large subunit (RPA1a) limits chiasma formation during rice meiosis[J]. Plant Physiology, 2021, 187(3): 1605-1618. doi: 10.1093/plphys/kiab365
[31] FERNANDES J B, DUHAMEL M, SEGUéLA -ARNAUD M, et al. FIGL1 and its novel partner FLIP form a conserved complex that regulates homologous recombination[J]. PLoS Genetics, 2018, 14(4): e1007317. doi: 10.1371/journal.pgen.1007317.
[32] DA INES O, ABE K, GOUBELY C, et al. Differing requirements for RAD51 and DMC1 in meiotic pairing of centromeres and chromosome arms in Arabidopsis thaliana[J]. PLoS Genetics, 2012, 8(4): e1002636. doi: 10.1371/journal.pgen.1002636.
[33] DA INES O, DEGROOTE F, GOUBELY C, et al. Meiotic recombination in Arabidopsis is catalysed by DMC1, with RAD51 playing a supporting role[J]. PLoS Genetics, 2013, 9(9): e1003787. doi: 10.1371/journal.pgen.1003787.
[34] CLOUD V, CHAN Y L, GRUBB J, et al. Rad51 is an accessory factor for Dmc1-mediated joint molecule formation during meiosis[J]. Science, 2012, 337(6099): 1222-1225. doi: 10.1126/science.1219379
[35] SEELIGER K, DUKOWIC-SCHULZE S, WURZ-WILDERSINN R, et al. BRCA2 is a mediator of RAD51- and DMC1-facilitated homologous recombination in Arabidopsis thaliana[J]. The New Phytologist, 2012, 193(2): 364-375. doi: 10.1111/j.1469-8137.2011.03947.x
[36] LIN Z, KONG H, NEI M, et al. Origins and evolution of the recA/RAD51 gene family: Evidence for ancient gene duplication and endosymbiotic gene transfer[J]. Proceedings of the National Academy of Sciences of the United States of America, 2006, 103(27): 10328-10333. doi: 10.1073/pnas.0604232103
[37] MERCIER R, MÉZARD C, JENCZEWSKI E, et al. The molecular biology of meiosis in plants[J]. Annual Review of Plant Biology, 2015, 66: 297-327. doi: 10.1146/annurev-arplant-050213-035923
[38] SU H, CHENG Z, HUANG J, et al. Arabidopsis RAD51, RAD51C and XRCC3 proteins form a complex and facilitate RAD51 localization on chromosomes for meiotic recombination[J]. PLoS Genetics, 2017, 13(5): e1006827. doi: 10.1371/journal.pgen.1006827.
[39] WANG Y, XIAO R, WANG H, et al. The Arabidopsis RAD51 paralogs RAD51B, RAD51D and XRCC2 play partially redundant roles in somatic DNA repair and gene regulation[J]. New Phytologist, 2014, 201(1): 292-304. doi: 10.1111/nph.12498
[40] DA INES O, DEGROOTE F, AMIARD S, et al. Effects of XRCC2 and RAD51B mutations on somatic and meiotic recombination in Arabidopsis thaliana[J]. The Plant Journal, 2013, 74(6): 959-970. doi: 10.1111/tpj.12182
[41] COLLINS I, NEWLON C S. Chromosomal DNA replication initiates at the same origins in meiosis and mitosis[J]. Molecular and Cellular Biology, 1994, 14(5): 3524-3534.
[42] SZOSTAK J W, ORR-WEAVER T L, ROTHSTEIN R J, et al. The double-strand-break repair model for recombination[J]. Cell, 1983, 33(1): 25-35. doi: 10.1016/0092-8674(83)90331-8
[43] TERASAWA M, OGAWA H, TSUKAMOTO Y, et al. Meiotic recombination-related DNA synthesis and its implications for cross-over and non-cross-over recombinant formation[J]. Proceedings of the National Academy of Sciences of the United States of America, 2007, 104(14): 5965-5970. doi: 10.1073/pnas.0611490104
[44] LU P L, HAN X W, QI J, et al. Analysis of Arabidopsis genome-wide variations before and after meiosis and meiotic recombination by resequencing Landsberg erecta and all four products of a single meiosis[J]. Genome Research, 2012, 22(3): 508-518. doi: 10.1101/gr.127522.111
[45] YANG H X, LU P L, WANG Y X, et al. The transcriptome landscape of Arabidopsis male meiocytes from high-throughput sequencing: The complexity and evolution of the meiotic process[J]. The Plant Journal, 2011, 65(4): 503-16. doi: 10.1111/j.1365-313X.2010.04439.x
[46] WANG Y X, CHENG Z H, HUANG J Y, et al. The DNA replication factor RFC1 is required for interference-sensitive meiotic crossovers in Arabidopsis thaliana[J]. PLoS Genetics, 2012, 8(11): e1003039. doi: 10.1371/journal.pgen.1003039.
[47] HUANG J Y, CHENG Z H, WANG C, et al. Formation of interference-sensitive meiotic cross-overs requires sufficient DNA leading-strand elongation[J]. Proceedings of the National Academy of Sciences of the United States of America, 2015, 112(40): 12534-12539. doi: 10.1073/pnas.1507165112
[48] HUANG J Y, COPENHAVER G P, MA H, et al. New insights into the role of DNA synthesis in meiotic recombination[J]. Science Bulletin, 2016, 61(16): 1260-1269. doi: 10.1007/s11434-016-1126-7
[49] WANG C, HUANG J Y, ZHANG J, et al. The largest subunit of DNA polymerase delta is required for normal formation of meiotic type I crossovers[J]. Plant Physiology, 2019, 179(2): 446-459. doi: 10.1104/pp.18.00861
[50] MCVEY M, KHODAVERDIAN V Y, MEYER D, et al. Eukaryotic DNA polymerases in homologous recombination[J]. Annual Review of Genetics, 2016, 50: 393-421. doi: 10.1146/annurev-genet-120215-035243
[51] MAGA G, STUCKI M, SPADARI S, et al. DNA polymerase switching: I: Replication factor C displaces DNA polymerase alpha prior to PCNA loading[J]. Journal of Molecular Biology, 2000, 295(4): 791-801. doi: 10.1006/jmbi.1999.3394
[52] DE MAAGD R A, LOONEN A, CHOUAREF J, et al. CRISPR/Cas inactivation of RECQ4 increases homeologous crossovers in an interspecific tomato hybrid[J]. Plant Biotechnology Journal, 2020, 18(3): 805-813. doi: 10.1111/pbi.13248
[53] MALOISEL L, BHARGAVA J, ROEDER G S. A role for DNA polymerase delta in gene conversion and crossing over during meiosis in Saccharomyces cerevisiae[J]. Genetics, 2004, 167(3): 1133-1142. doi: 10.1534/genetics.104.026260
[54] BISHOP D K, ZICKLER D. Early decision: Meiotic crossover interference prior to stable strand exchange and synapsis[J]. Cell, 2004, 117(1): 9-15. doi: 10.1016/S0092-8674(04)00297-1
[55] KURZBAUER M T, UANSCHOU C, CHEN D, et al. The recombinases DMC1 and RAD51 are functionally and spatially separated during meiosis in Arabidopsis[J]. The Plant Cell, 2012, 24(5): 2058-2070. doi: 10.1105/tpc.112.098459
[56] REN L J, ZHAO T T, ZHAO Y Z, et al. The E3 ubiquitin ligase DESYNAPSIS1 regulates synapsis and recombination in rice meiosis[J]. Cell Reports, 2021, 37(5): 109941. doi: 10.1016/j.celrep.2021.109941.
[57] BENYAHYA F, NADAUD I, DA INES O, et al. SPO11.2 is essential for programmed double-strand break formation during meiosis in bread wheat (Triticum aestivum L. )[J]. The Plant Journal, 2020, 104(1): 30-43. doi: 10.1111/tpj.14903
[58] CRISMANI W, GIRARD C, FROGER N, et al. FANCM limits meiotic crossovers[J]. Science, 2012, 336(6088): 1588-1590. doi: 10.1126/science.1220381
[59] GIRARD C, CRISMANI W, FROGER N, et al. FANCM-associated proteins MHF1 and MHF2, but not the other Fanconi anemia factors, limit meiotic crossovers[J]. Nucleic Acids Research, 2014, 42(14): 9087-9095. doi: 10.1093/nar/gku614
[60] FERNANDES J B, SÉGUÉLA-ARNAUD M, LARCHEVÊQUE C, et al. Unleashing meiotic crossovers in hybrid plants[J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(10): 2431-2436. doi: 10.1073/pnas.1713078114
[61] MERCIER R, JOLIVET S, VEZON D, et al. Two meiotic crossover classes cohabit in Arabidopsis: One is dependent on MER3, whereas the other one is not[J]. Current Biology, 2005, 15(8): 692-701. doi: 10.1016/j.cub.2005.02.056
[62] LYNN A, SOUCEK R, BÖRNER G V. ZMM proteins during meiosis: Crossover artists at work[J]. Chromosome Research, 2007, 15(5): 591-605. doi: 10.1007/s10577-007-1150-1
[63] MACAISNE N, VIGNARD J, MERCIER R. SHOC1 and PTD form an XPF-ERCC1-like complex that is required for formation of class I crossovers [J]. Journal of Cell Science, 2011, 124(Pt 16): 2687-91.
[64] CHELYSHEVA L, VEZON D, CHAMBON A, et al. The Arabidopsis HEI10 is a new ZMM protein related to Zip3[J]. PLoS Genetics, 2012, 8(7): e1002799. doi: 10.1371/journal.pgen.1002799.
[65] CHELYSHEVA L, GENDROT G, VEZON D, et al. Zip4/Spo22 is required for class I CO formation but not for synapsis completion in Arabidopsis thaliana[J]. PLoS Genetics, 2007, 3(5): e83. doi: 10.1371/journal.pgen.0030083.
[66] HIGGINS J D, ARMSTRONG S J H, FRANKLIN F C, et al. The Arabidopsis MutS homolog AtMSH4 functions at an early step in recombination: Evidence for two classes of recombination in Arabidopsis[J]. Genes & Development, 2004, 18(20): 2557-2570.
[67] HIGGINS J D, VIGNARD J, MERCIER R, et al. AtMSH5 partners AtMSH4 in the class I meiotic crossover pathway in Arabidopsis thaliana, but is not required for synapsis[J]. The Plant Journal, 2008, 55(1): 28-39. doi: 10.1111/j.1365-313X.2008.03470.x
[68] MAZINA O M, MAZIN A V, NAKAGAWA T, et al. Saccharomyces cerevisiae Mer3 helicase stimulates 3'-5' heteroduplex extension by Rad51: Implications for crossover control in meiotic recombination[J]. Cell, 2004, 117(1): 47-56. doi: 10.1016/S0092-8674(04)00294-6
[69] SNOWDEN T, ACHARYA S, BUTZ C, et al. hMSH4-hMSH5 recognizes holliday junctions and forms a meiosis-specific sliding clamp that embraces homologous chromosomes[J]. Molecular Cell, 2004, 15(3): 437-451. doi: 10.1016/j.molcel.2004.06.040
[70] DE MUYT A, PYATNITSKAYA A, ANDRÉANI J, et al. A meiotic XPF-ERCC1-like complex recognizes joint molecule recombination intermediates to promote crossover formation[J]. Genes & Development, 2018, 32(3/4): 283-296.
[71] PYATNITSKAYA A, ANDREANI J, GUÉROIS R, et al. The Zip4 protein directly couples meiotic crossover formation to synaptonemal complex assembly[J]. Genes & Development, 2022, 36(1/2): 53-69.
[72] REYNOLDS A, QIAO H Y, YANG Y, et al. RNF212 is a dosage-sensitive regulator of crossing-over during mammalian meiosis[J]. Nature Genetics, 2013, 45(3): 269-278. doi: 10.1038/ng.2541
[73] TOBY G G, GHERRABY W, COLEMAN T R, et al. A novel RING finger protein, human enhancer of invasion 10, alters mitotic progression through regulation of cyclin B levels[J]. Molecular and Cellular Biology, 2003, 23(6): 2109-2122. doi: 10.1128/MCB.23.6.2109-2122.2003
[74] ZIOLKOWSKI P A, UNDERWOOD C J, LAMBING C, et al. Natural variation and dosage of the HEI10 meiotic E3 ligase control Arabidopsis crossover recombination[J]. Genes & Development, 2017, 31(3): 306-317.
[75] MORGAN C, FOZARD J A, HARTLEY M, et al. Diffusion-mediated HEI10 coarsening can explain meiotic crossover positioning in Arabidopsis[J]. Nature Communications, 2021, 12(1): 4674. doi: 10.1038/s41467-021-24827-w.
[76] KURZBAUER M T, PRADILLO M, KERZENDORFER C, et al. Arabidopsis thaliana FANCD2 promotes meiotic crossover formation[J]. The Plant Cell, 2018, 30(2): 415-428. doi: 10.1105/tpc.17.00745
[77] MORGAN T H. Heredity and sex[Z] //Columbia University Lectures. The jessup lectures, New York: Columbia University Press, 1913.
[78] CAPILLA-PÉREZ L, DURAND S, HUREL A, et al. The synaptonemal complex imposes crossover interference and heterochiasmy in Arabidopsis[J]. Proceedings of the National Academy of Sciences of the United States of America, 2021, 118(12): e2023613118. doi: 10.1073/pnas.2023613118.
[79] NONOMURA K I, NAKANO M, MURATA K, et al. An insertional mutation in the rice PAIR2 gene, the ortholog of Arabidopsis ASY1, results in a defect in homologous chromosome pairing during meiosis[J]. Molecular Genetics and Genomics, 2004, 271(2): 121-129. doi: 10.1007/s00438-003-0934-z
[80] YUAN W Y, LI X W, CHANG Y X, et al. Mutation of the rice gene PAIR3 results in lack of bivalent formation in meiosis[J]. The Plant Journal, 2009, 59(2): 303-315. doi: 10.1111/j.1365-313X.2009.03870.x
[81] FRANCE M G, ENDERLE J, RÖHRIG S, et al. ZYP1 is required for obligate cross-over formation and cross-over interference in Arabidopsis[J]. Proceedings of the National Academy of Sciences of the United States of America, 2021, 118(14): e2021671118. doi: 10.1073/pnas.2021671118.
[82] POCHON G, HENRY I M, YANG C, et al. The Arabidopsis Hop1 homolog ASY1 mediates cross-over assurance and interference[EB/OL]. BioRxiv, 2022: 46164616 (2022-03-17) [2022-08-20]. https://doi.org/10.1101/2022.03.17.484635.
[83] DURAND S, LIAN Q, JING J, et al. Dual control of meiotic crossover patterning[EB/OL]. BioRxiv, 2022: 491364 (2022-05-11) [2022-08-20]. https://doi.org/10.1101/2022.05.11.491364.
[84] ROG O, KÖHLER S, DERNBURG A F. The synaptonemal complex has liquid crystalline properties and spatially regulates meiotic recombination factors[J]. eLife, 2017, 6: e21455. doi: 10.7554/eLife.21455.
[85] ZHANG L Y, STAUFFER W, ZWICKER D, et al. Crossover patterning through kinase-regulated condensation and coarsening of recombination nodules[EB/OL]. BioRxiv, 2021: 457865 (2021-08-26) [2022-08-20]. https://doi.org/10.1101/2021.08.26.457865.
[86] MAYER K F X, WAUGH R, LANGRIDGE P, et al. A physical, genetic and functional sequence assembly of the barley genome[J]. Nature, 2012, 491(7426): 711-716. doi: 10.1038/nature11543
[87] APPELS R, EVERSOLE K, STEIN N, et al. Shifting the limits in wheat research and breeding using a fully annotated reference genome[J]. Science, 2018, 361(6403): eaar7191. doi: 10.1126/science.aar7191.
[88] LI X, LI L, YAN J B. Dissecting meiotic recombination based on tetrad analysis by single-microspore sequencing in maize[J]. Nature Communications, 2015, 6: 6648. doi: 10.1038/ncomms7648.
[89] CONSORTIUM T G. The tomato genome sequence provides insights into fleshy fruit evolution[J]. Nature, 2012, 485(7400): 635-641. doi: 10.1038/nature11119
[90] MCCONAUGHY S J. Recombination hotspots in soybean [Glycine max (L. ) Merr. ], in Agronomy and Horticulture Department[D]. Lincoln: University of Nebraska, 2022.
[91] SCHMUTZ J, CANNON S B, SCHLUETER J, et al. Genome sequence of the palaeopolyploid soybean[J]. Nature, 2010, 463(7278): 178-183. doi: 10.1038/nature08670
[92] BLARY A, JENCZEWSKI E. Manipulation of crossover frequency and distribution for plant breeding[J]. Theoretical and Applied Genetics, 2019, 132(3): 575-592. doi: 10.1007/s00122-018-3240-1
[93] HSU Y M, FALQUE M, MARTIN O C. Quantitative modelling of fine-scale variations in the Arabidopsis thaliana crossover landscape[J]. Quantitative Plant Biology, 2022, 3: e3. doi: 10.1017/qpb.2021.17.
[94] PRATTO F, BRICK K, CHENG G, et al. Meiotic recombination mirrors patterns of germline replication in mice and humans[J]. Cell, 2021, 184(16): 4251-4267. doi: 10.1016/j.cell.2021.06.025
[95] UNDERWOOD C J, CHOI K, LAMBING C, et al. Epigenetic activation of meiotic recombination near Arabidopsis thaliana centromeres via loss of H3K9me2 and non-CG DNA methylation[J]. Genome Research, 2018, 28(4): 519-531. doi: 10.1101/gr.227116.117
[96] CHOI K, ZHAO X H, TOCK A J, et al. Nucleosomes and DNA methylation shape meiotic DSB frequency in Arabidopsis thaliana transposons and gene regulatory regions[J]. Genome Research, 2018, 28(4): 532-546. doi: 10.1101/gr.225599.117
[97] LIAN Q C, SOLIER V, WALKEMEIER B, et al. The megabase-scale crossover landscape is largely independent of sequence divergence[J]. Nature Communications, 2022, 13(1): 3828. doi: 10.1038/s41467-022-31509-8.
[98] YELINA N E, LAMBING C, HARDCASTLE T J, et al. DNA methylation epigenetically silences crossover hot spots and controls chromosomal domains of meiotic recombination in Arabidopsis[J]. Genes & Development, 2015, 29(20): 2183-2202.
[99] CHOI K, ZHAO X, KELLY K A, et al. Arabidopsis meiotic crossover hot spots overlap with H2A. Z nucleosomes at gene promoters[J]. Nature Genetics, 2013, 45(11): 1327-1336. doi: 10.1038/ng.2766
[100] MARAND A P, JANSKY S H, ZHAO H N, et al. Meiotic crossovers are associated with open chromatin and enriched with Stowaway transposons in potato[J]. Genome Biology, 2017, 18: 203. doi: 10.1186/s13059-017-1326-8.
[101] KIANIAN P M A, WANG M, SIMONS K, et al. Highresolution crossover mapping reveals similarities and differences of male and female recombination in maize[J]. Nature Communications, 2018, 9: 2370. doi: 10.1038/s41467-018-04562-5.
[102] TOCK A J, HOLLAND D M, JIANG W, et al. Crossover-active regions of the wheat genome are distinguished by DMC1, the chromosome axis, H3K27me3, and signatures of adaptation[J]. Genome Research, 2021, 31(9): 1614-1628. doi: 10.1101/gr.273672.120
[103] YELINA N E, CHOI K, CHELYSHEVA L, et al. Epigenetic remodeling of meiotic crossover frequency in Arabidopsis thaliana DNA methyltransferase mutants[J]. PLoS Genetics, 2012, 8(8): e1002844. doi: 10.1371/journal.pgen.1002844.
[104] PONT C, LEROY T, SEIDEL M, et al. Tracing the ancestry of modern bread wheats[J]. Nature Genetics, 2019, 51(5): 905-911. doi: 10.1038/s41588-019-0393-z
[105] ROWAN B A, HEAVENS D, FEUERBORN T R, et al. An ultra high-density Arabidopsis thaliana crossover map that refines the influences of structural variation and epigenetic features[J]. Genetics, 2019, 213(3): 771-787. doi: 10.1534/genetics.119.302406
[106] MARTÍN A C, SHAW P, PHILLIPS D, et al. Licensing MLH1 sites for crossover during meiosis[J]. Nature Communications, 2014, 5: 4580. doi: 10.1038/ncomms5580.
[107] TAM S M, HAYS J B, CHETELAT R T. Effects of suppressing the DNA mismatch repair system on homeologous recombination in tomato[J]. Theoretical and Applied Genetics, 2011, 123(8): 1445-1458. doi: 10.1007/s00122-011-1679-4
[108] ZHANG L, PICKERING R, MURRAY B. Direct measurement of recombination frequency in interspecific hybrids between Hordeum vulgare and H. bulbosum using genomic in situ hybridization [J]. Heredity, 1999, 83 ( Pt 3): 304-309.
[109] BLACKWELL A R, DLUZEWSKA J, SZYMANSKA-LEJMAN M, et al. MSH2 shapes the meiotic crossover landscape in relation to interhomolog polymorphism in Arabidopsis[J]. The EMBO Journal, 2020, 39(21): e104858. doi: 10.15252/embj.2020104858.
[110] GOODWILLIE C, KALISZ S, ECKERT C G, The evolutionary Enigma of mixed mating systems in plants: Occurrence, theoretical explanations, and empirical evidence [J]. Annual Review of Ecology, Evolution, and Systematics, 2005, 36(1): 47-79.
[111] MONNAHAN P, KOLÁŘ F, BADUEL P, et al. Pervasive population genomic consequences of genome duplication in Arabidopsis arenosa[J]. Nature Ecology & Evolution, 2019, 3(3): 457-468.
[112] BAUDAT F, BUARD J, GREY C, et al. PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice[J]. Science, 2010, 327(5967): 836-840. doi: 10.1126/science.1183439
[113] SMAGULOVA F, GREGORETTI I V, BRICK K, et al. Genome-wide analysis reveals novel molecular features of mouse recombination hotspots[J]. Nature, 2011, 472(7343): 375-378. doi: 10.1038/nature09869
[114] SHILO S, MELAMED-BESSUDO C, DORONE Y, et al. DNA crossover motifs associated with epigenetic modifications delineate open chromatin regions in Arabidopsis[J]. The Plant Cell, 2015, 27(9): 2427-2436. doi: 10.1105/tpc.15.00391
[115] LIU S Z, YEH C T, JI T M, et al. Mu transposon insertion sites and meiotic recombination events co-localize with epigenetic marks for open chromatin across the maize genome[J]. PLoS Genetics, 2009, 5(11): e1000733. doi: 10.1371/journal.pgen.1000733.
[116] SAINTENAC C, FAURE S, REMAY A, et al. Variation in crossover rates across a 3-Mb contig of bread wheat (Triticum aestivum) reveals the presence of a meiotic recombination hotspot[J]. Chromosoma, 2011, 120(2): 185-198. doi: 10.1007/s00412-010-0302-9
[117] DEMIRCI S, VAN DIJK A D J, SANCHEZ PEREZ G, et al. Distribution, position and genomic characteristics of crossovers in tomato recombinant inbred lines derived from an interspecific cross between Solanum lycopersicum and Solanum pimpinellifolium[J]. The Plant Journal, 2017, 89(3): 554-564. doi: 10.1111/tpj.13406
[118] HENDERSON I R. Control of meiotic recombination frequency in plant genomes[J]. Current Opinion in Plant Biology, 2012, 15(5): 556-561. doi: 10.1016/j.pbi.2012.09.002
[119] KUO P, DA INES O, LAMBING C. Rewiring meiosis for crop improvement[J]. Frontiers in Plant Science, 2021, 12: 708948. doi: 10.3389/fpls.2021.708948.
[120] UNDERWOOD C J, MERCIER R. Engineering apomixis: Clonal seeds approaching the fields[J]. Annual Review of Plant Biology, 2022, 73: 201-225. doi: 10.1146/annurev-arplant-102720-013958
[121] BERCHOWITZ L E, FRANCIS K E, BEY A L, et al. The role of AtMUS81 in interference-insensitive crossovers in A. thaliana[J]. PLoS Genetics, 2007, 3(8): e132. doi: 10.1371/journal.pgen.0030132.
[122] HARTUNG F, SUER S, PUCHTA H. Two closely related RecQ helicases have antagonistic roles in homologous recombination and DNA repair in Arabidopsis thaliana[J]. Proceedings of the National Academy of Sciences of the United States of America, 2007, 104(47): 18836-18841. doi: 10.1073/pnas.0705998104
[123] SÉGUÉLA-ARNAUD M, CHOINARD S, LARCHEVÊQUE C, et al. RMI1 and TOP3α limit meiotic CO formation through their C-terminal domains[J]. Nucleic Acids Research, 2017, 45(4): 1860-1871.
[124] SÉGUÉLA-ARNAUD M, CRISMANI W, LARCHEVÊQUE C, et al. Multiple mechanisms limit meiotic crossovers: TOP3α and two BLM homologs antagonize crossovers in parallel to FANCM[J]. Proceedings of the National Academy of Sciences of the United States of America, 2015, 112(15): 4713-4718. doi: 10.1073/pnas.1423107112
[125] GIRARD C, CHELYSHEVA L, CHOINARD S, et al. Correction: AAA-ATPase FIDGETIN-LIKE 1 and helicase FANCM antagonize meiotic crossovers by distinct mechanisms[J]. PLoS Genetics, 2015, 11(9): e1005448. doi: 10.1371/journal.pgen.1005448.
[126] BLARY A, GONZALO A, EBER F, et al. FANCM limits meiotic crossovers in Brassica crops[J]. Frontiers in Plant Science, 2018, 9: 368. doi: 10.3389/fpls.2018.00368.
[127] DESJARDINS S D, SIMMONDS J, GUTERMAN I, et al. FANCM promotes class I interfering crossovers and suppresses class II non-interfering crossovers in wheat meiosis[J]. Nature Communications, 2022, 13(1): 3644. doi: 10.1038/s41467-022-31438-6.
[128] MIEULET D, AUBERT G, BRES C, et al. Unleashing meiotic crossovers in crops[J]. Nature Plants, 2018, 4(12): 1010-1016. doi: 10.1038/s41477-018-0311-x
[129] RAZ A, DAHAN-MEIR T, MELAMED-BESSUDO C, et al. Redistribution of meiotic crossovers along wheat chromosomes by virus-induced gene silencing[J]. Frontiers in Plant Science, 2021, 11: 635139. doi: 10.3389/fpls.2020.635139.
[130] LI X, YU M S, BOLAÑOS-VILLEGAS P, et al. Fanconi anemia ortholog FANCM regulates meiotic crossover distribution in plants[J]. Plant Physiology, 2021, 186(1): 344-360. doi: 10.1093/plphys/kiab061
[131] ARRIETA M, MACAULAY M, COLAS I, et al. An induced mutation in HvRECQL4 increases the overall recombination and restores fertility in a barley HvMLH3 mutant background[J]. Frontiers in Plant Science, 2021, 12: 706560. doi: 10.3389/fpls.2021.706560.
[132] DE MUYT A, ZHANG L R, PIOLOT T, et al. E3 ligase Hei10: A multifaceted structure-based signaling molecule with roles within and beyond meiosis[J]. Genes & Development, 2014, 28(10): 1111-1123.
[133] SERRA H, LAMBING C, GRIFFIN C H, et al. Massive crossover elevation via combination of HEI10 and recq4a recq4b during Arabidopsis meiosis[J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(10): 2437-2442. doi: 10.1073/pnas.1713071115
[134] TAAGEN E, BOGDANOVE A J, SORRELLS M E. Counting on crossovers: Controlled recombination for plant breeding[J]. Trends in Plant Science, 2020, 25(5): 455-465. doi: 10.1016/j.tplants.2019.12.017
[135] MELAMED-BESSUDO C, LEVY A A. Deficiency in DNA methylation increases meiotic crossover rates in euchromatic but not in heterochromatic regions in Arabidopsis[J]. Proceedings of the National Academy of Sciences of the United States of America, 2012, 109(16): E981-E988. doi: 10.1073/pnas.1120742109.
[136] LIU C L, CAO Y W, HUA Y F, et al. Concurrent disruption of genetic interference and increase of genetic recombination frequency in hybrid rice using CRISPR/Cas9[J]. Frontiers in Plant Science, 2021, 12: 757152. doi: 10.3389/fpls.2021.757152.
[137] WANG K J, WANG C, LIU Q, et al. Increasing the genetic recombination frequency by partial loss of function of the synaptonemal complex in rice[J]. Molecular Plant, 2015, 8(8): 1295-1298. doi: 10.1016/j.molp.2015.04.011
[138] NAGESWARAN D C, KIM J, LAMBING C, et al. HIGH CROSSOVER RATE1 encodes PROTEIN PHOSPHATASE X1 and restricts meiotic crossovers in Arabidopsis[J]. Nature Plants, 2021, 7(4): 452-467. doi: 10.1038/s41477-021-00889-y
[139] HOCHHOLDINGER F, BALDAUF J A. Heterosis in plants[J]. Current Biology, 2018, 28(18): R1089-R1092. doi: 10.1016/j.cub.2018.06.041
[140] SPILLANE C, CURTIS M D, GROSSNIKLAUS U. Apomixis technology development: Virgin births in farmers’ fields?[J]. Nature Biotechnology, 2004, 22(6): 687-691. doi: 10.1038/nbt976
[141] WANG C, LIU Q, SHEN Y, et al. Clonal seeds from hybrid rice by simultaneous genome engineering of meiosis and fertilization genes[J]. Nature Biotechnology, 2019, 37(3): 283-286. doi: 10.1038/s41587-018-0003-0
[142] MARIMUTHU M P A, JOLIVET S, RAVI M, et al. Synthetic clonal reproduction through seeds[J]. Science, 2011, 331(6019): 876. doi: 10.1126/science.1199682.
[143] D'ERFURTH I, JOLIVET S, FROGER N, et al. Turning meiosis into mitosis[J]. PLoS Biology, 2009, 7(6): e1000124. doi: 10.1371/journal.pbio.1000124.
[144] MIEULET D, JOLIVET S, RIVARD M, et al. Turning rice meiosis into mitosis[J]. Cell Research, 2016, 26(11): 1242-1254. doi: 10.1038/cr.2016.117
[145] CROMER L, HEYMAN J, TOUATI S, et al. OSD1 promotes meiotic progression via APC/C inhibition and forms a regulatory network with TDM and CYCA1;2/TAM[J]. PLoS Genetics, 2012, 8(7): e1002865. doi: 10.1371/journal.pgen.1002865.
[146] RAVI M, MARIMUTHU M P A, SIDDIQI I. Gamete formation without meiosis in Arabidopsis[J]. Nature, 2008, 451(7182): 1121-1124.
[147] HSU P D, LANDER E S, ZHANG F. Development and applications of CRISPR-Cas9 for genome engineering[J]. Cell, 2014, 157(6): 1262-1278. doi: 10.1016/j.cell.2014.05.010
[148] RÖNSPIES M, DORN A, SCHINDELE P, et al. CRISPR-Cas-mediated chromosome engineering for crop improvement and synthetic biology[J]. Nature Plants, 2021, 7(5): 566-573. doi: 10.1038/s41477-021-00910-4
[149] SARNO R, VICQ Y, UEMATSU N, et al. Programming sites of meiotic crossovers using Spo11 fusion proteins[J]. Nucleic Acids Research, 2017, 45(19): e164. doi: 10.1093/nar/gkx739.
[150] YELINA N E, HOLLAND D, GONZALEZ-JORGE S, et al. Coexpression of MEIOTIC-TOPOISOMERASE VIB-dCas9 with guide RNAs specific to a recombination hotspot is insufficient to increase crossover frequency in Arabidopsis[J]. G3 (Genes Genomes Genetics), 2022, 12(7): jkac105. doi: 10.1093/g3journal/jkac105.
[151] MANCERA E, BOURGON R, BROZZI A, et al. High-resolution mapping of meiotic crossovers and non-crossovers in yeast[J]. Nature, 2008, 454(7203): 479-485. doi: 10.1038/nature07135
[152] WANG Y Z, VAN RENGS W M J, ZAIDAN M W A M, et al. Meiosis in crops: From genes to genomes[J]. Journal of Experimental Botany, 2021, 72(18): 6091-6109. doi: 10.1093/jxb/erab217
[153] FILLER HAYUT S, MELAMED BESSUDO C, LEVY A A. Targeted recombination between homologous chromosomes for precise breeding in tomato[J]. Nature Communications, 2017, 8: 15605. doi: 10.1038/ncomms15605.
[154] KOURANOV A, ARMSTRONG C, SHRAWAT A, et al. Demonstration of targeted crossovers in hybrid maize using CRISPR technology[J]. Communications Biology, 2022, 5(1): 53. doi: 10.1038/s42003-022-03004-9.
[155] BEYING N, SCHMIDT C, PACHER M, et al. CRISPR-Cas9-mediated induction of heritable chromosomal translocations in Arabidopsis[J]. Nature Plants, 2020, 6(6): 638-645. doi: 10.1038/s41477-020-0663-x
[156] SCHMIDT C, FRANSZ P, RÖNSPIES M, et al. Changing local recombination patterns in Arabidopsis by CRISPR/Cas mediated chromosome engineering[J]. Nature Communications, 2020, 11(1): 4418. doi: 10.1038/s41467-020-18277-z.
[157] SCHWARTZ C, LENDERTS B, FEIGENBUTZ L, et al. CRISPR-Cas9-mediated 75.5-Mb inversion in maize[J]. Nature Plants, 2020, 6(12): 1427-1431. doi: 10.1038/s41477-020-00817-6
[158] GALLEGO-BARTOLOMÉ J, GARDINER J, LIU W L, et al. Targeted DNA demethylation of the Arabidopsis genome using the human TET1 catalytic domain[J]. Proceedings of the National Academy of Sciences of the United States of America, 2018, 115(9): E2125-E2134. doi: 10.1073/pnas.1716945115.
[159] GALLEGO-BARTOLOMÉ J, LIU W L, KUO P H, et al. Co-targeting RNA polymerases IV and V promotes efficient de novo DNA methylation in Arabidopsis[J]. Cell, 2019, 176(5): 1068-1082.e19. doi: 10.1016/j.cell.2019.01.029
[160] DIRKS R, VAN DUN K, DE SNOO C B, et al. Reverse breeding: A novel breeding approach based on engineered meiosis[J]. Plant Biotechnology Journal, 2009, 7(9): 837-845. doi: 10.1111/j.1467-7652.2009.00450.x
[161] WIJNKER E, DEURHOF L, VAN DE BELT J, et al. Hybrid recreation by reverse breeding in Arabidopsis thaliana[J]. Nature Protocols, 2014, 9(4): 761-772. doi: 10.1038/nprot.2014.049
[162] WIJNKER E, VAN DUN K, DE SNOO C B, et al. Reverse breeding in Arabidopsis thaliana generates homozygous parental lines from a heterozygous plant[J]. Nature Genetics, 2012, 44(4): 467-470. doi: 10.1038/ng.2203
[163] CALVO‐BALTANÁS V, WIJNEN C L, YANG C, et al. Meiotic crossover reduction by virus‐induced gene silencing enables the efficient generation of chromosome substitution lines and reverse breeding in Arabidopsis thaliana[J]. The Plant Journal, 2020, 104(5): 1437-1452. doi: 10.1111/tpj.14990
-
期刊类型引用(5)
1. 詹鹏麟,张扬,林建新,许静,庄炜,卢和顶,陈山虎,廖长见. 闽双色6号籽粒发育中叶酸合成代谢基因挖掘与分析. 中国细胞生物学学报. 2025(02): 165-176 .  百度学术
百度学术
2. 谢梓明,聂笑一. 农作物基因组学数据库研究进展. 农业装备与车辆工程. 2025(03): 151-156 .  百度学术
百度学术
3. 李健雄,陈建酉,涂从勇,朱素素,江远汉,何文杰,陈荣彬,何秀英. 试验基地建设新模式的探索与实践——以广东省农业科学院水稻研究所试验基地为例. 中国稻米. 2024(05): 136-140 .  百度学术
百度学术
4. 朱庆莹. 水稻机械化作业对农业生产效率的影响. 农机使用与维修. 2024(10): 31-34 .  百度学术
百度学术
5. 李健雄,姜敏,朱素素,郑智华,吴平波,陈煬杨,黄章慧,涂从勇,张陆军,陈建酉. 新时期农业科研单位开源节流工作的实践与思考——以广东省农业科学院水稻研究所为例. 农业科技管理. 2024(06): 103-107 .  百度学术
百度学术
其他类型引用(0)


 下载:
下载: