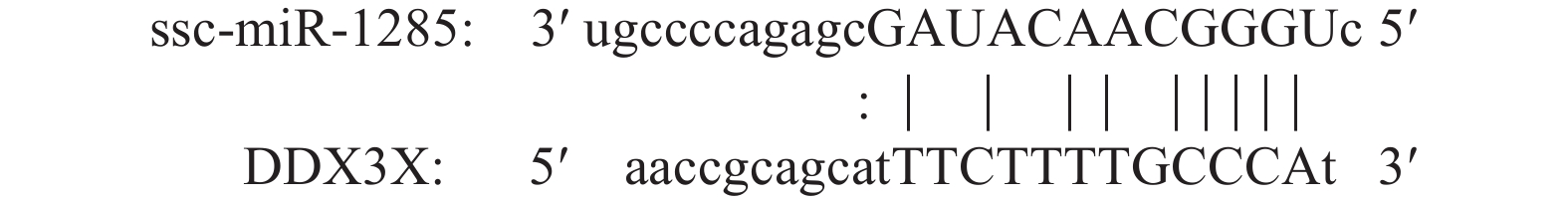Regulation effects of microRNA-1285 and its target DDX3X on Senecavirus A infected PK-15 cells
-
摘要:目的
探究MicroRNA-1285(miR-1285)及其靶标DDX3X在猪塞内卡病毒(Senecavirus A,SVA)感染PK-15细胞中的调控作用。
方法利用qRT-PCR、双荧光素酶活性及Western blot等方法研究miR-1285和DDX3X对I型干扰素(IFN-β)分泌及RIG-I信号通路的作用,分析miR-1285及DDX3X对SVA 3C蛋白基因表达的影响。
结果SVA感染PK-15细胞后,miR-1285表达量显著升高,并且miR-1285与DDX3X存在负靶向关系,二者可促进IFN-β转录及蛋白水平的表达。miR-1285通过靶向DDX3X对RIG-I信号通中的MAVS、TRAF3信号分子起调控作用。对于SVA 3C蛋白基因,DDX3X可以显著抑制其转录,并且可以逆转miR-1285所诱导的上调趋势。
结论SVA感染PK-15细胞后,宿主miR-1285及其靶标DDX3X对IFN-β及病毒3C蛋白的表达具有调控作用,研究结果将为明确miRNAs调控SVA感染的分子机制奠定基础,并为SVA的防控和诊断提供新的科学依据。
-
关键词:
- MicroRNA-1285 /
- DDX3X /
- 猪塞内卡病毒 /
- IFN-β /
- RIG-I信号通路
Abstract:ObjectiveTo explore the regulation roles of microRNA-1285 (miR-1285) and its target DDX3X in Senecavirus A (SVA) infected PK-15 cells.
MethodBy qRT-PCR, double luciferase activity and Western blot, the effects of miR-1285 and its target DDX3X on IFN-β secretion and the RIG-I signaling pathway were studied, and their effects on the expression of SVA 3C protein gene were analyzed.
ResultIn SVA infected PK-15 cells, the expression of miR-1285 increased significantly, and there was a negative targeting relationship between miR-1285 and DDX3X. Both miR-1285 and DDX3X promoted the transcription and protein expression of IFN-β. MiR-1285 regulated MAVS and TRAF3 signaling molecules in the RIG-I signaling pathway by targeting DDX3X. For SVA 3C protein, DDX3X significantly inhibited the transcription of 3C and reversed the up-regulation trend induced by miR-1285.
ConclusionAfter infecting PK-15 cells with SVA, host miR-1285 and its target DDX3X can regulate the expression of IFN-β and the viral 3C protein, which will lay a foundation for clarifying the molecular mechanism of miRNAs regulating SVA infection, and provide a new scientific basis for the prevention, control and diagnosis of SVA.
-
Keywords:
- MiRNA-1285 /
- DDX3X /
- Senecavirus A /
- IFN-β /
- RIG-I signal pathway
-
-
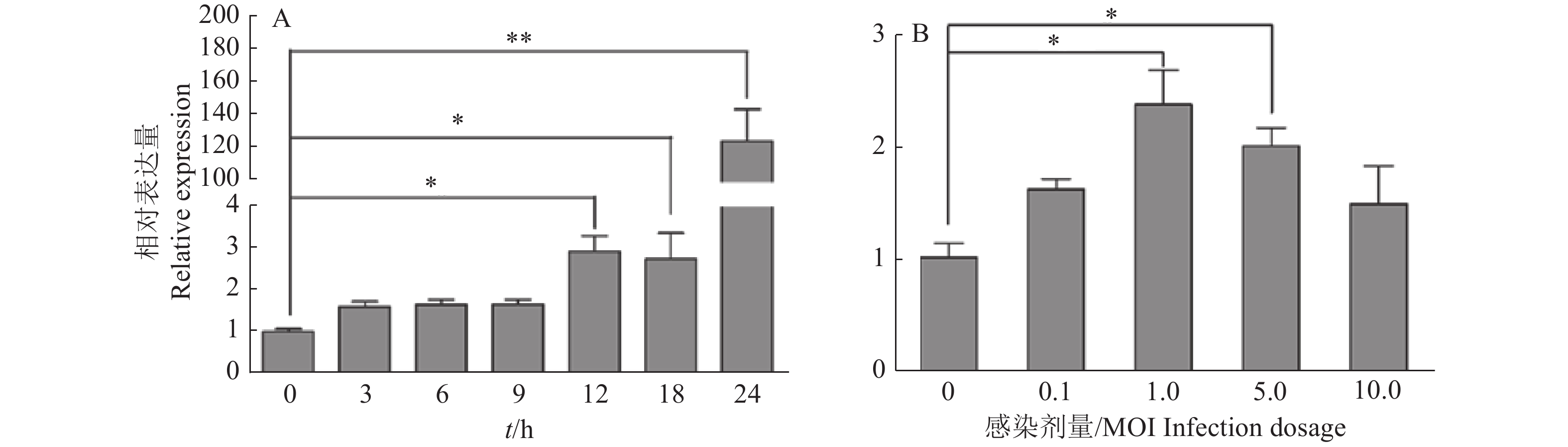
图 1 不同SVA感染时间(A)和感染剂量(B)条件下PK-15细胞中miR-1285的表达量
“*”“**”分别表示处理与对照在P < 0.05和P < 0.01水平差异显著(Duncan’s法)
Figure 1. Expression of miR-1285 in PK-15 cells infected by SVA at different infection time (A) and dosages (B)
“*” and “**” represented statistical difference in comparison with control group at P < 0.05 and P < 0.01 levels respectively (Duncan’s method)
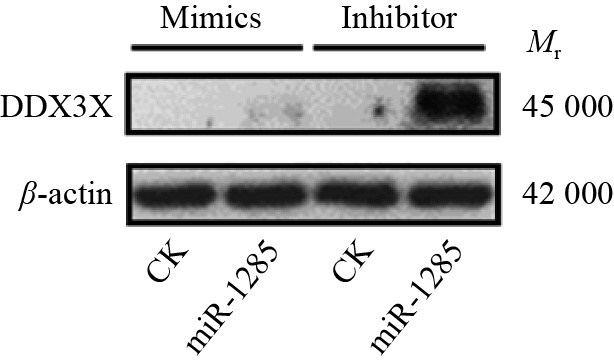
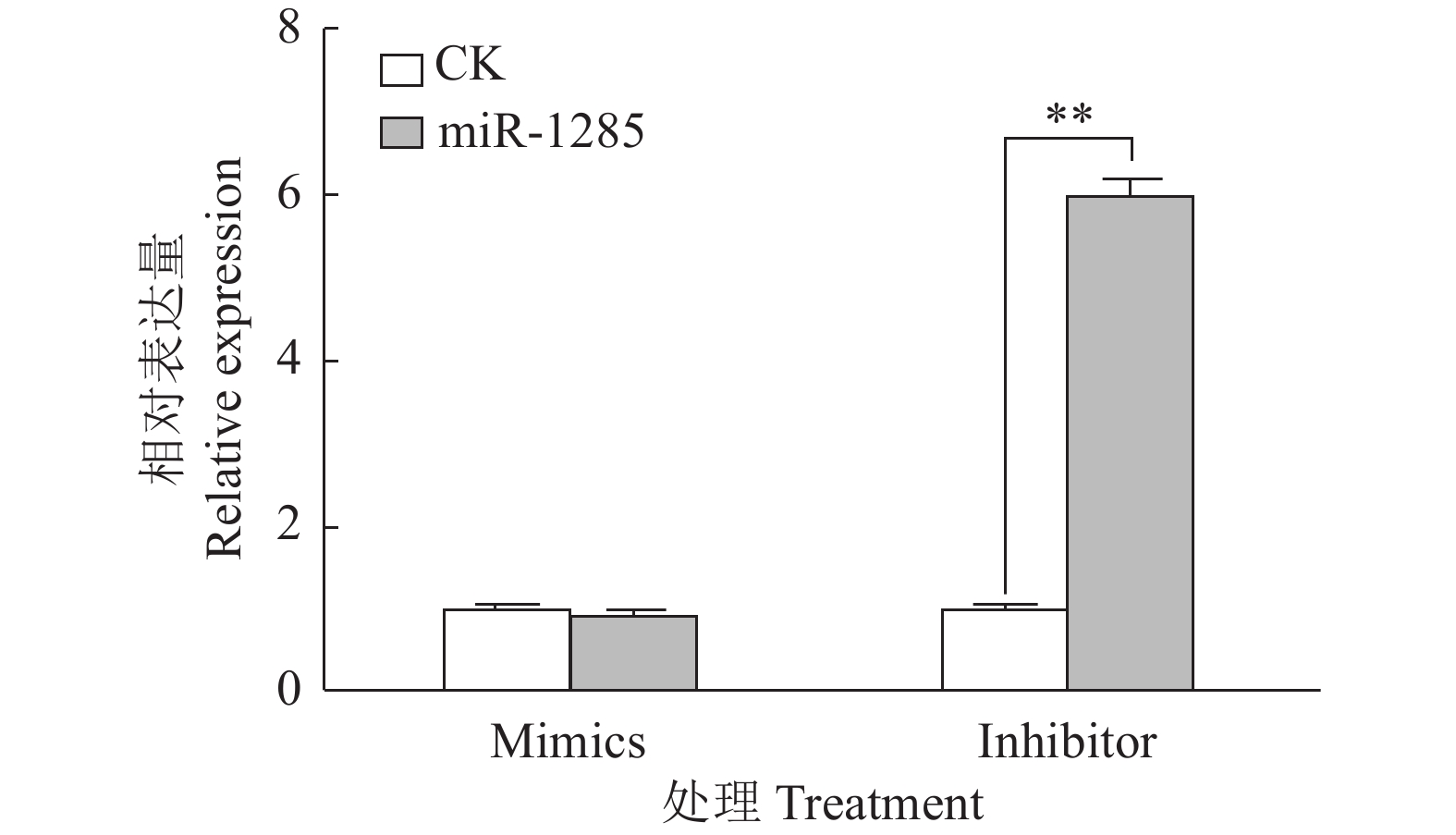
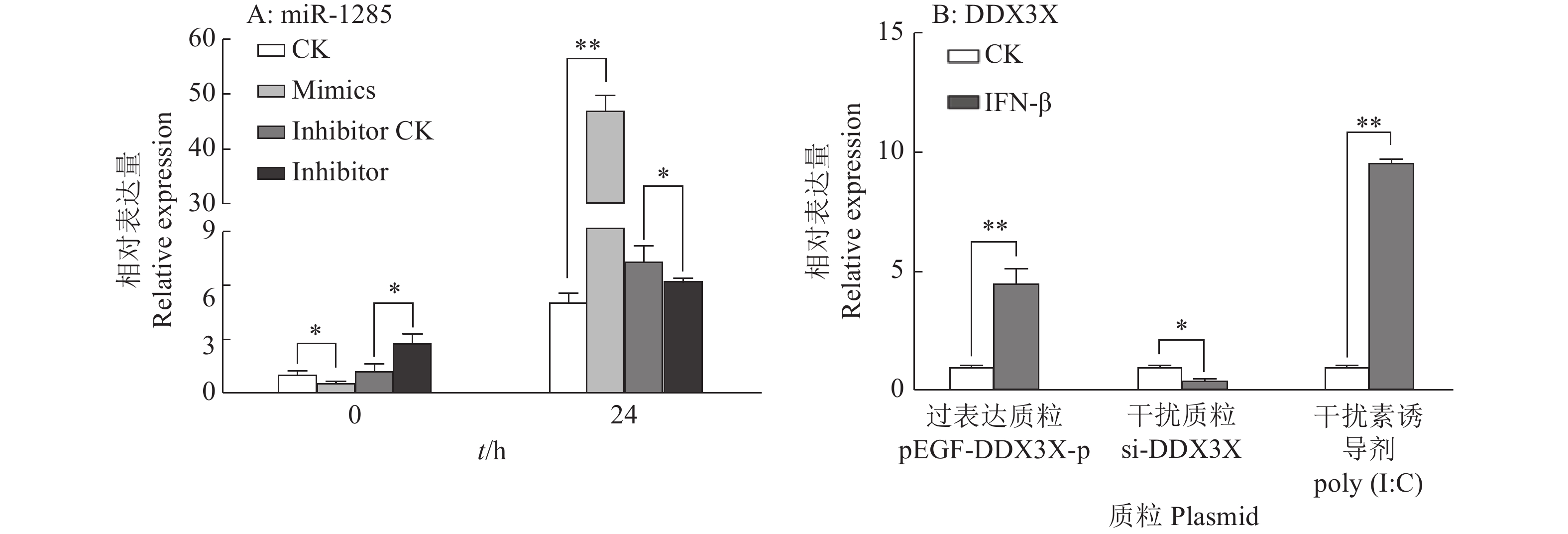
图 4 转染miR-1285 mimics、inhibitor至PK-15细胞后DDX3X的mRNA相对表达量
“**”表示处理与对照在P < 0.01水平差异显著 (Duncan’s 法)
Figure 4. The relative expression of DDX3X mRNA after transfection of miR-1285 mimics and inhibitor into PK-15 cells
“**” represented statistical difference in comparison with control group at P < 0.01 level (Duncan’s method)
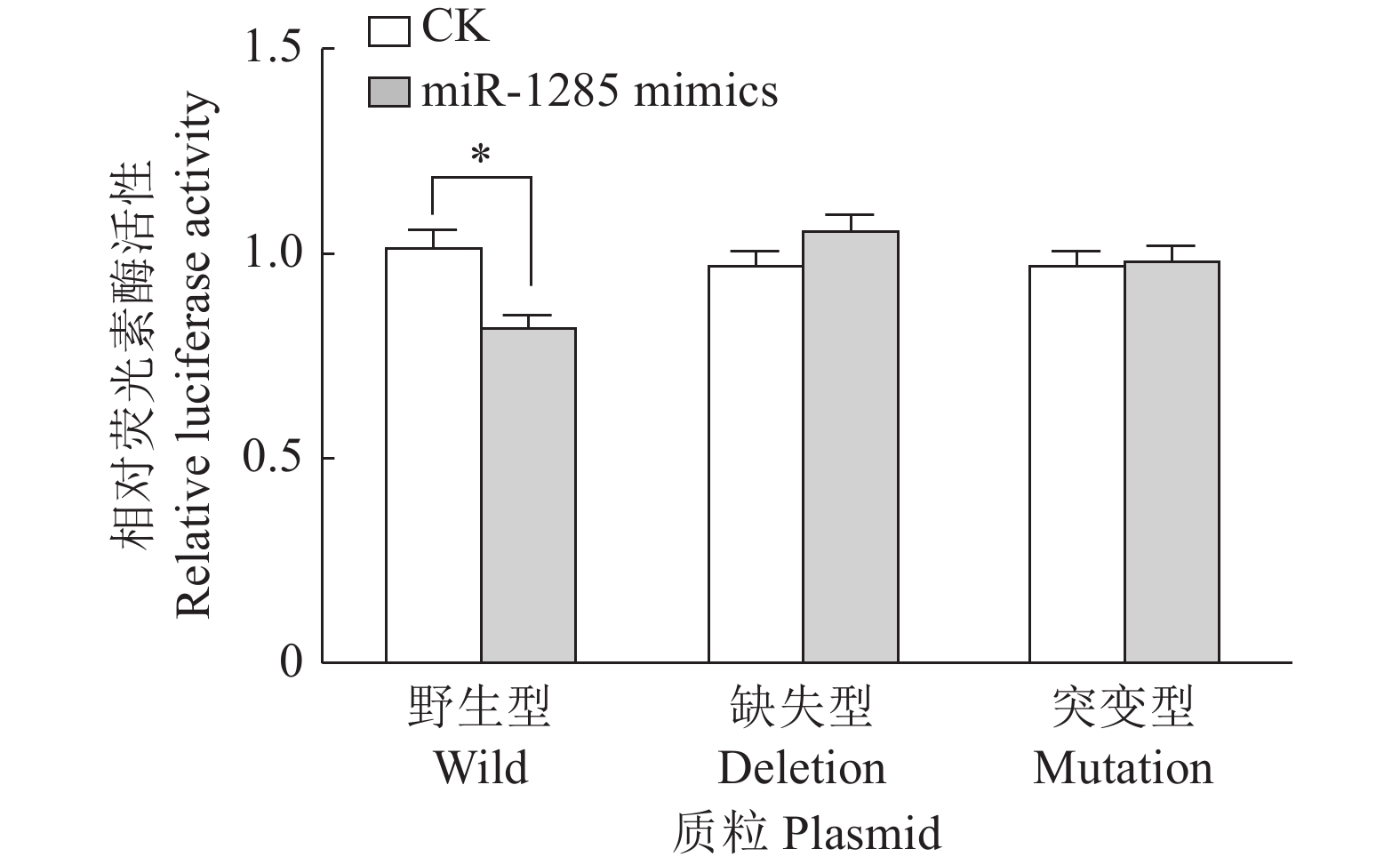
图 5 转染不同DDX3X重组载体质粒至PK-15细胞后miR-1285双荧光素酶活性
“*”表示处理与对照在P < 0.05水平差异显著 (Duncan’s 法)
Figure 5. The relative dual-luciferase activity of miR-1285 after transfection of different DDX3X recombinant vector plasmids into PK-15 cells
“*” represented statistical difference in comparison with control group at P < 0.05 level (Duncan’s method)
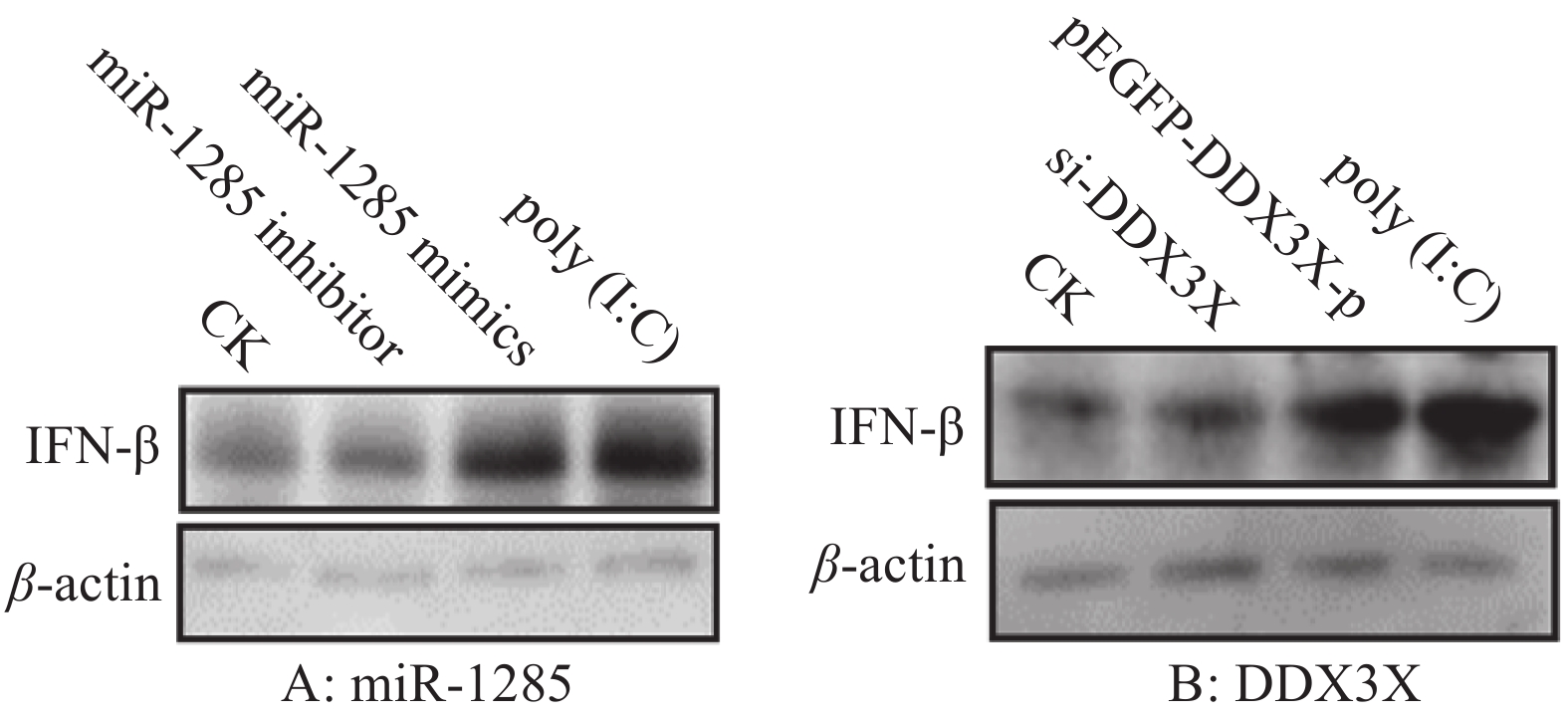
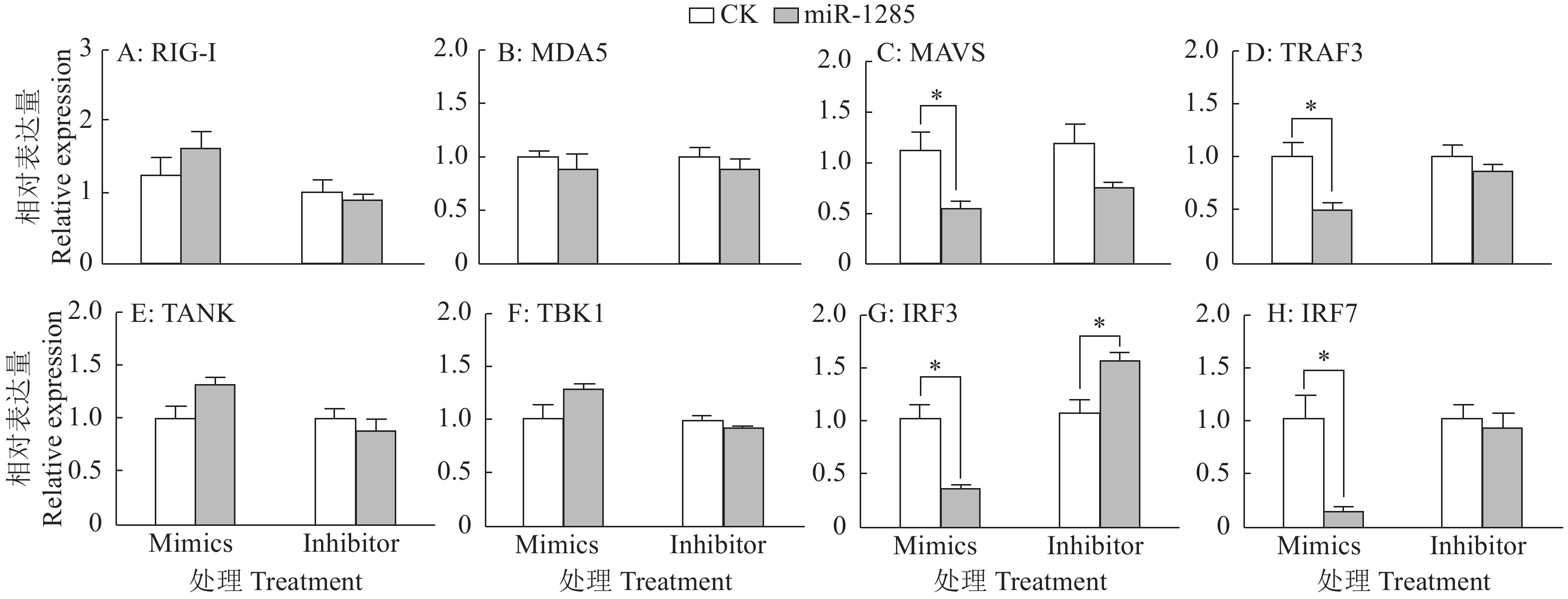
图 6 miR-1285及其靶标DDX3X对IFN-β mRNA表达的调控作用
“*”和“**”分别表示处理与对照在P < 0.05和P < 0.01水平差异显著(Duncan’s法)
Figure 6. Regulation effects of miR-1285 and its target DDX3X on the mRNA expression of IFN-β
“*” and “**” represented statistical difference in comparison with control group at P < 0.05 and P < 0.01 levels respectively (Duncan’s method)
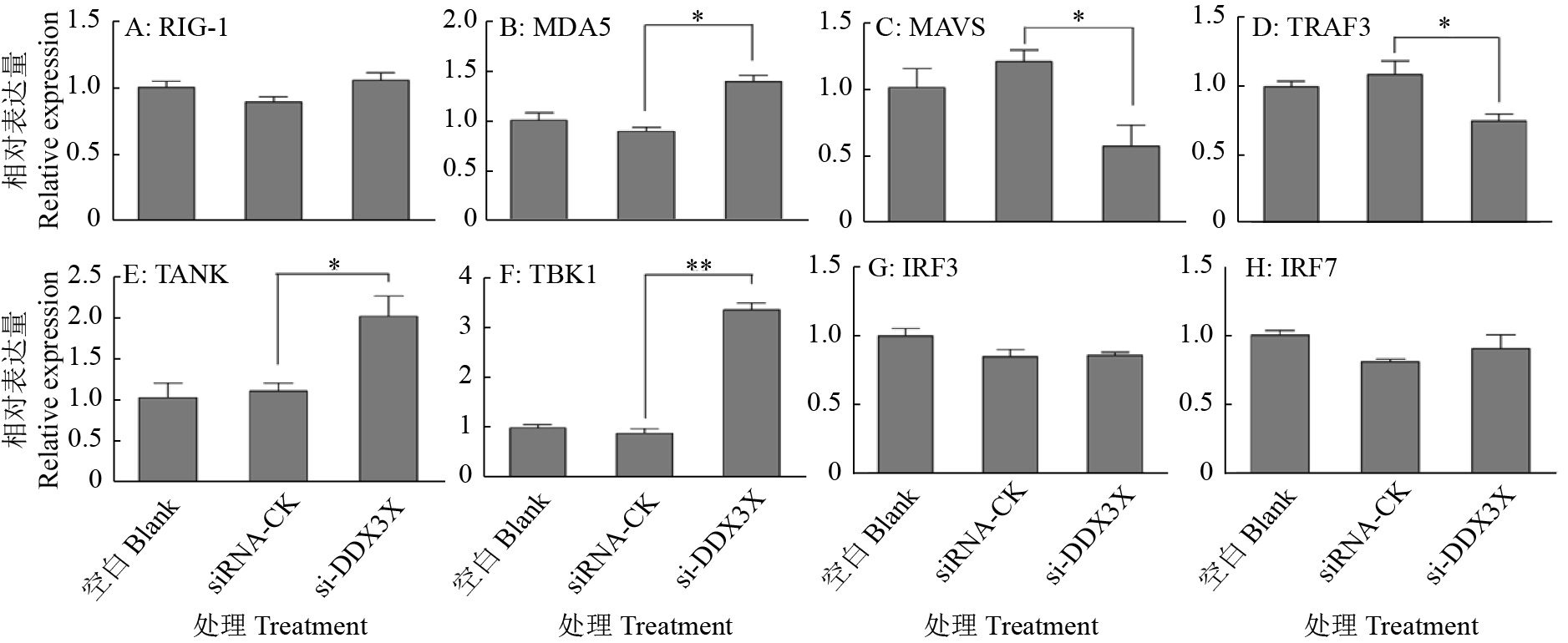
图 9 DDX3X沉默对RIG-I通路信号转导分子的影响
“*”和“**”分别表示处理与对照在P < 0.05和P < 0.01水平差异显著(Duncan’s法)
Figure 9. Effects of DDX3X silencing on signal transduction molecules of the RIG-I pathway
“*” and “**” represented statistical difference in comparison with control group at P < 0.05 and P < 0.01 levels respectively (Duncan’s method)
表 1 基因引物序列
Table 1 Primer sequences of genes
基因名称 Gene name 引物/探针序列(5′→3′) Primer/Probe sequence θ退火/ ℃ Annealing temperature 产物大小/bp Product size 文献 Reference RIG-I F: ATCCCAGCAACGAGAA 60 188 [36] R: GCCACGTCCAGTCAAT MDA5 F: GAGGAATCAGCACGAGGAA 58 73 [37] R: GTCAGTAATCCACTGGGA MAVS F: ATAGCCAGCCTTTCTCGG 60 237 [36] R: TAGCCTCAGTCTTGACCTCTTC TRAF3 F: GTGTCAAGAAGGCATCG 60 164 [36] R: CCTCAAACTGGCAATCA TANK F: GGACGCCTTGAACTACCTGT 60 119 R: GCCTGCCGAAAGGCTTCATA TBK1 F: GCCTTTCTCGGGGTCTTCAA 60 74 R: ACACTTTTCCTGATCCGCCT IRF3 F: CCAGTGGTGCCTACACTCCT 61 191 [38] R: AGAGGTGTCTGGCTCAGGAA IRF7 F: CGCCTCCTGGAAAACCAA 60 76 [37] R: CCCTGAGTTGTCCTGCAACA IFN-β F: GCTAACAAGTGCATCCTCCAAA 60 77 [39] R:AGCACATCATAGCTCATGGAAAGA GAPDH F: ACATGGCCTCCAAGGAGTAAGA 60 106 [40] R: GATCGAGTTGGGGCTGTGACT SVA-3C F: GAGCTTCAATCTCCTAGA 59 115 R: GTGTCATCATTCTCGTTAG 探针 Probe CAGACATTCGAGCCAAGCAACAA 69 -
[1] YATES L A, NORBURY C J, GIBERT R J C. The long and short of microRNA[J]. Cell, 2013, 153(3): 516-519. doi: 10.1016/j.cell.2013.04.003
[2] CHENG J, WU R, LONG L, et al. MiRNA-451a targets IFN regulatory factor 8 for the progression of systemic lupus erythematosus[J]. Inflammation, 2017, 40(2): 676-687. doi: 10.1007/s10753-017-0514-8
[3] WANG L, ZHOU L, HU D, et al. Porcine reproductive and respiratory syndrome virus suppresses post-transcriptionally the protein expression of IFN-β by upregulating cellular microRNAs in porcine alveolar macrophages in vitro[J]. Experimental and Therapeutic Medicine, 2018, 15(1): 115-126.
[4] MORIN R D, O’CONNOR M D, GRIFFITH M, et al. Application of massively parallel sequencing to microRNA profiling and discovery in human embryonic stem cells[J]. Genome Research, 2008, 18(4): 610-621. doi: 10.1101/gr.7179508
[5] ANTHON C, TAFER H, HAVGAARD J H, et al. Structured RNAs and synteny regions in the pig genome[J]. BMC Genomics, 2014, 15: 459. doi: 10.1186/1471-2164-15-459.
[6] GAO X, WANG Y, ZHAO H, et al. Plasma miR-324-3p and miR-1285 as diagnostic and prognostic biomarkers for early stage lung squamous cell carcinoma[J]. Oncotarget, 2016, 7(37): 59664-59675. doi: 10.18632/oncotarget.11198
[7] BORRELLI N, DENARO M, UGOLINI C, et al. MiRNA expression profiling of ‘noninvasive follicular thyroid neoplasms with papillary-like nuclear features’ compared with adenomas and infiltrative follicular variants of papillary thyroid carcinomas[J]. Modern Pathology, 2017, 30(1): 39-51. doi: 10.1038/modpathol.2016.157
[8] LIU J, YAN J, ZHOU C, et al. MiR-1285-3p acts as a potential tumor suppressor miRNA via down regulating JUN expression in hepatocellular carcinoma[J]. Tumor Biology, 2015, 36: 219-225. doi: 10.1007/s13277-014-2622-5
[9] HUANG H, XIONG G, SHEN P, et al. MicroRNA-1285 inhibits malignant biological behaviors of human pancreatic cancer cells by negative regulation of YAP1[J]. Neoplasma, 2017, 64(3): 358-366. doi: 10.4149/neo_2017_306
[10] LIAO J, LI Q, HU Z, et al. Mitochondrial miR-1285 regulates copper-induced mitochondrial dysfunction and mitophagy by impairing IDH2 in pig jejunal epithelial cells[J]. Journal of Hazardous Materials, 2022, 422: 126899. doi: 10.1016/j.jhazmat.2021.126899.
[11] VENKATARAMANNAN S, GADEK M, CALVIELLO L, et al. DDX3X and DDX3Y are redundant in protein synthesis[J]. RNA, 2021, 27(12): 1577-1588. doi: 10.1261/rna.078926.121
[12] VAN VOSS M R H, KAMMERS K, VESUNA F, et al. Global effects of DDX3 inhibition on cell cycle regulation identified by a combined phosphoproteomics and single cell tracking approach[J]. Translational Oncology, 2018, 11(3): 755-763. doi: 10.1016/j.tranon.2018.04.001
[13] HEATON S M, BORG N A, DIXIT V M. Ubiquitin in the activation and attenuation of innate antiviral immunity[J]. Journal of Experimental Medicine, 2016, 213(1): 1-13. doi: 10.1084/jem.20151531
[14] HATHAICHOTI S, VISITNONTHACHAI D, NGAMSIRI P, et al. Paraquat induces extrinsic pathway of apoptosis in A549 cells by induction of DR5 and repression of anti-apoptotic proteins, DDX3 and GSK3 expression[J]. Toxicology in Vitro, 2017, 42: 123-129. doi: 10.1016/j.tiv.2017.04.016
[15] LIN T C. DDX3X multifunctionally modulates tumor progression and serves as a prognostic indicator to predict cancer outcomes[J]. International Journal of Molecular Sciences, 2019, 21(1): 281. doi: 10.3390/ijms21010281.
[16] SOULAT D, BŰRCKSTŰMMER T, WESTERMAYER S, et al. The DEAD-box helicase DDX3X is a critical component of the TANK-binding kinase 1-dependent innate immune response[J]. EMBO Journal, 2008, 27(15): 2135-2146. doi: 10.1038/emboj.2008.126
[17] SCHRŐDER M, BARAN M, BOWIE A G. Viral targeting of DEAD box protein 3 reveals its role in TBK1/IKKepsilon-mediated IRF activation[J]. EMBO Journal, 2008, 27(15): 2147-2157. doi: 10.1038/emboj.2008.143
[18] OSHIUMI H, SAKAI K, MATSUMOT M, et al. DEAD/H BOX 3 (DDX3) helicase binds the RIG-I adaptor IPS-1 to up-regulate IFN-β-inducing potential[J]. European Journal of Immunology, 2010, 40(4): 940-948. doi: 10.1002/eji.200940203
[19] SZAPPANOS D, TSCHISMAROV R, PERLOT T, et al. The RNA helicase DDX3X is an essential mediator of innate antimicrobial immunity[J]. PLoS Pathogens, 2018, 14(11): e1007397. doi: 10.1371/journal.ppat.1007397
[20] ADAMS M J, LEFKOWITZ E J, KING A M Q, et al. Ratification vote on taxonomic proposals to the international committee on taxonomy of viruses[J]. Archives of Virology, 2016, 161(10): 2921-2949. doi: 10.1007/s00705-016-2977-6
[21] VANNUCCI F A, LINHARES D C L, BARCELLOS D E S N, et al. Identification and complete genome of Seneca Valley virus in vesicular fluid and sera of pigs affected with idiopathic vesicular disease, Brazil[J]. Transboundary Emerging Disease, 2015, 62: 589-593. doi: 10.1111/tbed.12410
[22] CANNING P, CANON A, BATES J L, et al. Neonatal mortality, vesicular lesions and lameness associated with Senecavirus A in a U. S. sow farm[J]. Transboundary and Emerging Disease, 2016, 63(4): 373-378. doi: 10.1111/tbed.12516
[23] ZHANG X L, ZHU Z X, YANG F, et al. Review of Seneca Valley virus: A call for increased surveillance and research[J]. Frontiers in Microbiology, 2018, 9: 940. doi: 10.3389/fmicb.2018.00940.
[24] FERNANDES M H V, MAGGIOLI M F, OTTA J, et al. Senecavirus A 3C protease mediates host cell apoptosis late in infection[J]. Frontiers in Immunology, 2019, 10: 363. doi: 10.3389/fimmu.2019.00363.
[25] LEME R A, OLIVEIRA T E S, ALFIERI A F, et al. Pathological, immunohistochemical and molecular findings associated with Senecavirus A-induced lesions in neonatal piglets[J]. Journal of Comparative Pathology, 2016, 155(2/3): 145-155.
[26] HAUSE B M, MYERS O, DUFF J, et al. Senecavirus A in pigs, United States, 2015[J]. Emerging Infectious Diseases, 2016, 22(7): 1323-1325. doi: 10.3201/eid2207.151591
[27] XU W, HOLE K, GOOLIA M, et al. Genome wide analysis of the evolution of Senecavirus A from swine clinical material and assembly yard environmental samples[J]. PLoS One, 2017, 12(5): e0176964. doi: 10.1371/journal.pone.0176964
[28] WU Q, ZHAO X, BAI Y, et al. The first identification and complete genome of Senecavirus A affecting pig with idiopathic vesicular disease in China[J]. Transboundary and Emerging Disease, 2017, 64(5): 1633-1640. doi: 10.1111/tbed.12557
[29] SAENG-CLUTO K, RODTIAN P, TEMEEYASEN G, et al. The first detection of Senecavirus A in pigs in Thailand, 2016[J]. Transboundary and Emerging Disease, 2018, 65(1): 285-288. doi: 10.1111/tbed.12654
[30] SUN D, VANNUCCI F, KNUTSON T P, et al. Emergence and whole-genome sequence of Senecavirus A in Colombia[J]. Transboundary and Emerging Diseases, 2017, 64(5): 1346-1349. doi: 10.1111/tbed.12669
[31] ARZT J, BERTRAM M R, VU L T, et al. First detection and genome sequence of Senecavirus A in Vietnam[J]. Microbiology Resource Announcements, 2019, 8(3). doi: 10.1128/MRA.01247-18.
[32] SUN Y, CHENG J, WU R T, et al. Phylogenetic and genome analysis of 17 novel Senecavirus A isolates in Guangdong Province, 2017[J]. Frontiers in Veterinary Science, 2018, 5: 314. doi: 10.3389/fvets.2018.00314.
[33] QIAN S, FAN W, QIAN P, et al. Isolation and full-genome sequencing of Seneca Valley virus in piglets from China, 2016[J]. Virology Journal, 2016, 13: 173. doi: 10.1186/s12985-016-0631-2.
[34] ZHU Z, YANG F, CHEN P, et al. Emergence of novel Seneca Valley virus strains in China, 2017[J]. Transboundary and Emerging Diseases, 2017, 64(4): 1024-1029. doi: 10.1111/tbed.12662
[35] WANG H, LI C, ZHAO B, et al. Complete genome sequence and phylogenetic analysis of Senecavirus A isolated in Northeast China in 2016[J]. Archives of Virology, 2017, 162(10): 3173-3176. doi: 10.1007/s00705-017-3480-4
[36] ZHANG J, MIAO J, HOU J, et al. The effects of H3N2 swine influenza virus infection on TLRs and RLRs signaling pathways in porcine alveolar macrophages[J]. Virology Journal, 2015, 12: 61. doi: 10.1186/s12985-015-0284-6.
[37] 谢立兰. 伪狂犬病毒和猪传染性胃肠炎病毒诱导β干扰素产生的分子机制研究[D]. 武汉: 华中农业大学, 2011. [38] ISLAM M A, GROßE-BRINKHAUS C, PRŐLL M J, et al. Deciphering transcriptome profiles of peripheral blood mononuclear cells in response to PRRSV vaccination in pigs[J]. BMC Genomics, 2016, 17: 641. doi: 10.1186/s12864-016-2849-1.
[39] 王荡. 口蹄疫病毒Lpro和3Cpro调控宿主抗病毒天然免疫反应的分子机制研究[D]. 武汉: 华中农业大学, 2011. [40] 王国庆. 口蹄疫病毒2B蛋白拮抗RIG-I抗病毒作用研究[D]. 兰州: 甘肃农业大学, 2015. [41] DERRICK T, ROBERTS C H, RAJASEKHAR M, et al. Conjunctival microRNA expression in inflammatory trachomatous scarring[J]. PLoS Neglected Tropical Diseases, 2013, 7(3): e2117. doi: 10.1371/journal.pntd.0002117
[42] KAUR S, KRISHN S R, RACHAGANI S, et al. Significance of microRNA-based biomarkers for pancreatic cancer[J]. Annals of Translational Medicine, 2015, 3(18): 277.
[43] LIU Y, XU X, XU X, et al. MicroRNA-193a-3p inhibits cell proliferation in prostate cancer by targeting cyclin D1[J]. Oncology Letters, 2017, 14(5): 5121-5128.
[44] MORGUL M H, KLUNK S, ANASTASIADOU Z, et al. Diagnosis of HCC for patients with cirrhosis using miRNA profiles of the tumor-surrounding tissue: A statistical model based on stepwise penalized logistic regression[J]. Experimental and Molecular Pathology, 2016, 101(2): 165-171. doi: 10.1016/j.yexmp.2016.07.014
[45] POIRIER J T, DOBROMILSKAVA I, MORIARTY W F, et al. Selective tropism of Seneca Valley virus for variant subtype small cell lung cancer[J]. Journal of the National Cancer Institute, 2013, 105(14): 1059-1065. doi: 10.1093/jnci/djt130
[46] ZHOU Z H, SUN Y, YAN X L, et al. Swine acute diarrhea syndrome coronavirus (SADS-CoV) antagonizes interferon-β production via blocking IPS-1 and RIG-I[J]. Virus Research, 2020, 278: 197843. doi: 10.1016/j.virusres.2019.197843.
[47] ANDREJEVA J, CHILDS K S, YOUNG D F, et al. The V proteins of paramyxoviruses bind the IFN-inducible RNA helicase, mda-5, and inhibit its activation of the IFN-beta promoter[J]. Proceedings of the National Academy of Sciences of the United States of America, 2004, 101(49): 17264-17269. doi: 10.1073/pnas.0407639101
[48] CAO L, GE X, GAO Y, et al. Porcine epidemic diarrhea virus inhibits dsRNA-induced interferon-beta production in porcine intestinal epithelial cells by blockade of the RIG-I-mediated pathway[J]. Virology Journal, 2015, 12: 127. doi: 10.1186/s12985-015-0345-x.
[49] CHEN Q, LIU Q, LIU D, et al. Molecular cloning, functional characterization and antiviral activity of porcine DDX3X[J]. Biochemical and Biophysical Research Communications, 2014, 443(4): 1169-1175. doi: 10.1016/j.bbrc.2013.12.098
[50] ZHAO X, WU Q, BAI Y, et al. Phylogenetic and genome analysis of seven Senecavirus A isolates in China[J]. Transboundary and Emerging Diseases, 2017, 64(6): 2075-2082. doi: 10.1111/tbed.12619
[51] THOMPSON S R, SARNOW P. Regulation of host cell translation by viruses and effects on cell function[J]. Current Opinion in Microbiology, 2000, 3(4): 366-370. doi: 10.1016/S1369-5274(00)00106-5
[52] QIAN S, FAN W, LIU T, et al. Seneca Valley virus suppresses host type I interferon production by targeting adaptor proteins MAVS, TRIF, and TANK for cleavage[J]. Journal of Virology, 2017, 91(16): e00823-17.
[53] LIU T, LI X, WU M, et al. Seneca Valley virus 2C and 3Cpro induce apoptosis via mitochondrion-mediated intrinsic pathway[J]. Frontiers in Microbiology, 2019, 10: 1202. doi: 10.3389/fmicb.2019.01202.
[54] XUE Q, LIU H, ZHU Z, et al. Seneca Valley Virus 3C protease negatively regulates the type I interferon pathway by acting as a viral deubiquitinase[J]. Antiviral Research, 2018, 160: 183-189. doi: 10.1016/j.antiviral.2018.10.028


 下载:
下载: