Analyses of drug resistance, multi-locus sequence typing and genetic evolution of Peudomonas aeruginosa in pheasant farms
-
摘要:目的
探究广东省七彩山鸡养殖场中铜绿假单胞菌Pseudomonas aeruginosa的流行特点、耐药性、多位点序列分型(Multi-locus sequence typing,MLST)及遗传进化背景,为临床合理用药提供参考。
方法从广东省3个规模化七彩山鸡养殖场收集孵化死胚及周围环境样本进行铜绿假单胞菌的分离鉴定,采用K-B纸片扩散法测试其对22种抗菌药物的敏感性,利用MLST分析铜绿假单胞菌分离株的分子流行特点。将各ST型的7个管家基因序列按顺序拼接,用MEGA7软件对拼接好的序列进行遗传进化分析。
结果在采集的514份样本(死胚样本405份、环境样本109份)中共分离到铜绿假单胞菌145株(分离率28.2%),其中,24株来源于环境样本(分离率22.0%,24/109),121株来源于死胚样本(分离率29.9%,121/405)。145株铜绿假单胞菌除对氨苄西林、卡那霉素和萘啶酸天然耐药外,对复方新诺明、氯霉素和四环素的耐药性较强,耐药率分别为100%、80.0%和77.2%,其次为头孢噻肟(耐药率23.4%)。除天然耐药外,多重耐药的铜绿假单胞菌占比达73.1%(106/145),且对亚胺培南也显现出耐药性。MLST结果显示,89株耐药谱较广的铜绿假单胞菌可分为18个ST型,其中,6个ST型是本研究新发现的。铜绿假单胞菌死胚分离株的优势ST型是ST-260,环境分离株以ST-2100和ST-3202为主。遗传进化分析表明,环境和死胚样品中的铜绿假单胞菌具有较高的相似性。
结论本研究调查的3个七彩山鸡养殖场的死胚及周围环境中均存在不同程度的铜绿假单胞菌感染或污染,且分离株具有较强的耐药性;建议在养殖过程中不仅要加强饲养管理、提高生物安全意识,还应根据药敏试验结果合理选择抗菌药。
Abstract:ObjectiveTo investigate the epidemic characteristics, drug resistance, multi-locus sequence typing (MLST) and genetic evolution background of Pseudomonas aeruginosa in Guangdong Province, and provide a reference for clinical rational drug use.
MethodSamples from dead embryos of pheasant and their surrounding environment were collected for separation and identification of P. aeruginosa. The K-B paper disk diffusion method was used to analyze P. aeruginosa sensitivity to 22 kinds of antimicrobials. The MLST method was applied to analyze the molecular epidemiology of P. aeruginosa strains. Seven house-keeping genes of each ST type were spliced sequentially, and we used MEGA7 software to conduct genetic evolution analysis to the spliced sequence.
ResultA total of 145 P. aeruginosa strains (isolation rate 28.2%) were isolated from the collected 514 samples (405 dead embryo samples and 109 environment samples), including 24 strains from environmental samples (isolation rate 22.0%, 24/109) and 121 strains from dead embryos (isolation rate 29.9%, 121/405). Antibiotic drug sensitivity test showed that 145 strains of P. aeruginosa were naturally resistant to ampicillin, kanamycin and nalidixic acid, strongly resistant to complex sulfamethoxazole, chloramphenicol, tetracycline, followed by cefotaxime, with the drug resistance rates of 100%, 80.0%, 77.2% and 23.4% respectively. Except the natural drug resistance, the proportion of multiple drug resistant P. aeruginosa was up to 73.1% (106/145), and a certain proportion of P. aeruginosa strains which were resistant to imipenem appeared. MLST analysis showed that 89 P. aeruginosa strains with a broad spectrum of resistance were divided into 18 ST types, presenting high diversity. Among them, six ST types were the new types discovered in this study. The dead embryo isolates of P. aeruginosa were mainly ST-260, and the environmental sample isolates were mainly ST-2100 and ST-3202. Genetic evolution analysis showed that P. aeruginosa strains in environment were closely related to those in dead embryos.
ConclusionThere are different degrees of P. aeruginosa infection or contamination in dead embryos and surrounding environment of three pheasant farms, and the isolates have strong drug resistance. Therefore, it is recommended that we should not only strengthen the breeding management and raise the awareness of bio-safety in the process of breeding, but also use antibacterial drugs reasonably according to the results of the drug sensitivity test.
-
-
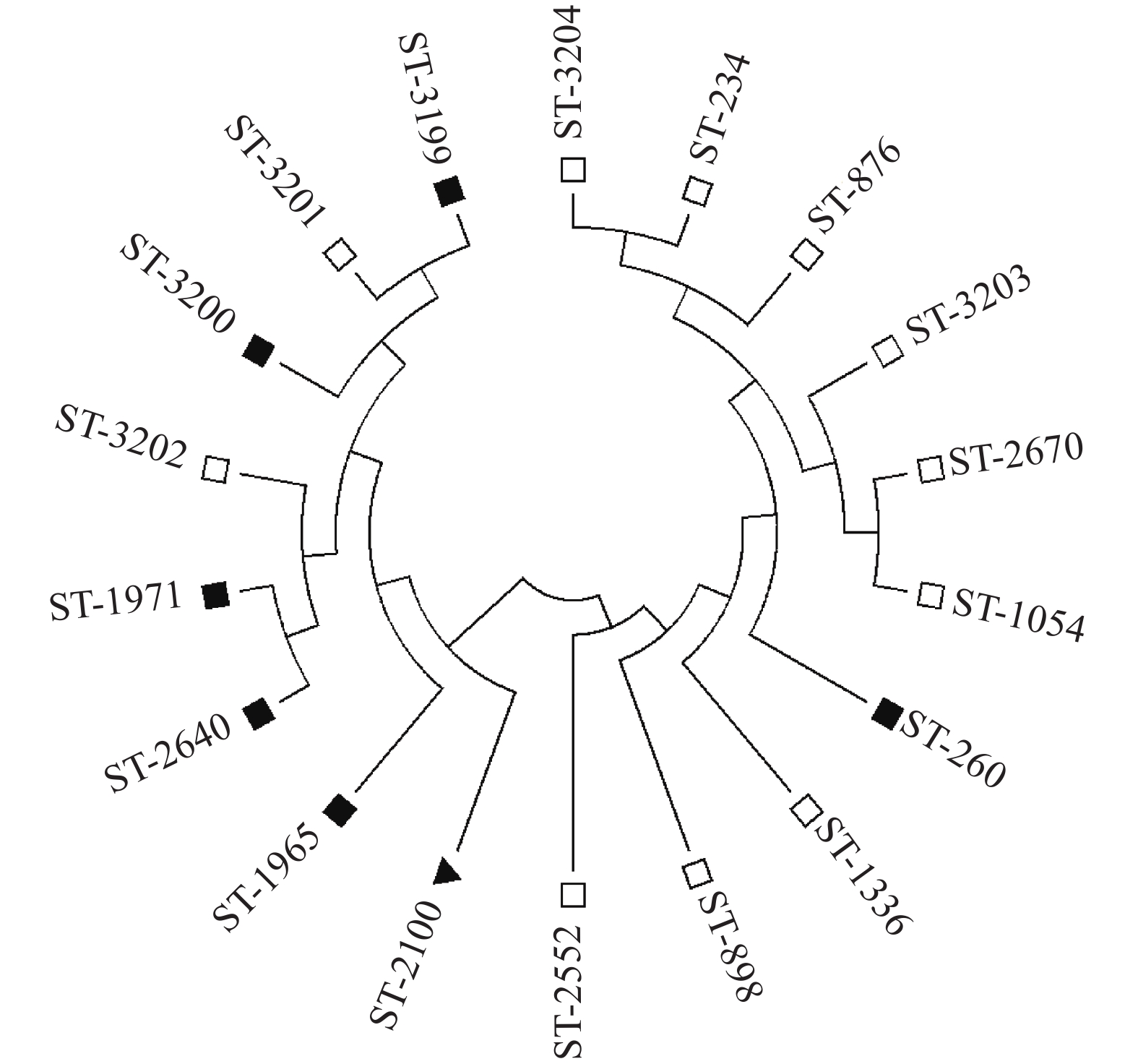
图 3 死胚和环境样品中不同铜绿假单胞菌ST型的遗传进化分析
“■”:铜绿假单胞菌死胚分离株所属的ST型;“□”:铜绿假单胞菌环境样品分离株所属的ST型;“▲”:2种样品共有的ST型
Figure 3. Genetic evolution analysis of different Pseudomonas aeruginosa ST types isolated from dead embryo and environment samples
“■”: ST type of Pseudomonas aeruginosa strain isolated from dead embryo; “□”: ST type of P. aeruginosa strain isolated from environment sample; “▲”: The common ST type of P. aeruginosa strain isolated from dead embryo and environment sample
表 1 不同七彩山鸡养殖场中铜绿假单胞菌临床分离株的数量及分离率
Table 1 Quantity and isolation rate of clinical isolates of Pseudomonas aeruginosa from different breeding poultry farms
养殖场
Poultry
farm样本类型
Sample type样品采集数
Collected sample number阳性样品数
Positive sample number分离率/%
Isolation rateA 死胚
Dead embryo222 85 38.3
(87/227)环境
Environment5 2 B 死胚
Dead embryo96 24 21.2
(28/132)环境
Environment36 4 C 死胚
Dead embryo87 12 19.4
(30/155)环境
Environment68 18 合计
Total514 145 28.2
(145/514)表 2 死胚和环境样品中耐药铜绿假单胞菌菌株数量及占比1)
Table 2 Number and proportion of drug-resistant Pseudomonas aeruginosa strain in dead embryo and environment samples
抗菌药
Antimicrobial死胚
Dead embryo环境
Environment总计
TotalAMP 121(100.0) 24(100.0) 145(100.0) NA 121(100.0) 24(100.0) 145(100.0) K 121(100.0) 24(100.0) 145(100.0) SXT 121(100.0) 24(100.0) 145(100.0) C 92(76.0) 24(100.0) 116(80.0) TE 90(74.4) 22(91.7) 112(77.2) CTX 27(22.3) 7(29.2) 34(23.4) LEV 7(5.8) 11(45.8) 18(12.4) CIP 7(5.8) 9(37.5) 16(11.1) TOB 7(5.8) 6(25.0) 13(9.0) CN 8(6.6) 5(20.8) 13(9.0) PRL 10(8.3) — 10(6.9) CFP 7(5.8) 3(12.5) 10(6.9) ATM 7(5.8) 1(4.2) 8(5.5) CAZ 5(4.1) — 5(3.5) FEP 4(3.3) — 4(2.8) TZP 3(1.9) — 3(2.1) IPM 1(0.8) 1(4.2) 2(1.4) MER — — — AMI — — — CES — — — POB — — — 1) AMP:氨苄西林,SXT:复方新诺明,NA:萘啶酸,K:卡那霉素,TE:四环素,C:氯霉素,PRL:哌拉西林,CTX:头孢噻肟,FEP:头孢吡肟,CAZ:头孢他啶,CFP:头孢哌酮,IPM:亚胺培南,ATM:氨曲南,TZP:哌拉西林/他唑巴坦,TOB:妥布霉素,CN:庆大霉素,CIP:环丙沙星,LEV:左氧氟沙星,IPM:亚胺培南,MER:美罗培南,AMI:阿米卡星,CES:头孢哌酮/舒巴坦,POB:多黏菌素B;括号内数据为占比;“—”表示未检测到耐药菌株
1) AMP: Ampicillin, SXT: Complex sulfamethoxazole, NA: Nalidixic acid, K: Kanamycin, TE: Tetracycline, C: Chloramphenicol, PRL: Piperacillin, CTX: Cefotaxime, FEP: Cefepime, CAZ: Ceftazidime, CFP: Cefoperazone, IPM: Imipenem, ATM: Aztreonam, TZP: Piperacillin/tazobactam, TOB: Tobramycin, CN: Gentamicin, CIP: Ciprofloxacin, LEV: Levofloxacin, IPM: Imipenem, MER: Meropenem, AMI: Amikacin, CES: Cefoperazone/sulbactam, POB: Polymyxin B; The data in brackets are proportions; “—” indicates no drug-resistant isolate表 3 铜绿假单胞菌的耐药谱及对应菌株数量
Table 3 Drug resistance spectrum and strain quantity of Pseudomonas aeruginosa
耐药数
Drug-resistant number耐药谱1)
Drug resistance spectrum菌株数量
Strain number4 AMP+K+NA+SXT 14 5 AMP+K+TE+NA+SXT 7 AMP+K+C+NA+SXT 18 6 AMP+K+TE+C+NA+SXT 59 AMP+CTX+K+TE+NA+SXT 6 7 AMP+CTX+K+TE+C+NA+SXT 12 AMP+PRL+CTX+CFP+K+NA+SXT 1 AMP+K+TE+C+NA+LEV+SXT 3 8 AMP+K+TE+C+NA+CIP+LEV+SXT 5 AMP+CTX+CFP+K+TE+C+NA+SXT 1 AMP+CTX+ATM+K+TE+C+NA+SXT 1 9 AMP+PRL+K+TOB+CN+TE+C+NA+SXT 2 10 AMP+PRL+CTX+K+TOB+CN+TE+C+NA+SXT 1 AMP+K+TOB+CN+TE+C+NA+CIP+LEV+SXT 2 AMP+CTX+K+CN+TE+C+NA+CIP+LEV+SXT 1 AMP+CTX+K+TOB+CN+TE+C+NA+CIP+SXT 1 AMP+CFP+K+TOB+TE+C+NA+CIP+LEV+SXT 1 11 AMP+PRL+CTX+CAZ+CFP+ATM+K+TE+C+NA+SXT 1 AMP+PRL+CTX+FEP+CAZ+CFP+ATM+K+TE+NA+SXT 1 AMP+CTX+K+TOB+CN+TE+C+NA+CIP+LEV+SXT 3 13 AMP+PRL+CTX+FEP+CAZ+CFP+ATM+TZP+K+TE+C+NA+SXT 3 AMP+CTX+IPM+ATM+K+TOB+CN+TE+C+NA+CIP+LEV+SXT 1 15 AMP+PRL+CTX+CFP+IPM+ATM+K+TOB+CN+TE+C+NA+CIP+LEV+SXT 1 1)AMP:氨苄西林,SXT:复方新诺明,NA:萘啶酸,K:卡那霉素,TE:四环素,C:氯霉素,PRL:哌拉西林,CTX:头孢噻肟,FEP:头孢吡肟,CAZ:头孢他啶,CFP:头孢哌酮,IPM:亚胺培南,ATM:氨曲南,TZP:哌拉西林/他唑巴坦,TOB:妥布霉素,CN:庆大霉素,CIP:环丙沙星,LEV:左氧氟沙星
1) AMP: Ampicillin, SXT: Complex sulfamethoxazole, NA: Nalidixic acid, K: Kanamycin, TE: Tetracycline, C: Chloramphenicol, PRL: Piperacillin, CTX: Cefotaxime, FEP: Cefepime, CAZ: Ceftazidime, CFP: Cefoperazone, IPM: Imipenem, ATM: Aztreonam, TZP: Piperacillin/tazobactam, TOB: Tobramycin, CN: Gentamicin, CIP: Ciprofloxacin, LEV: Levofloxacin表 4 89株铜绿假单胞菌菌株各ST型菌株数量及来源
Table 4 Quantity and source of each ST type in 89 Pseudomonas aeruginosa strains
ST型1)
ST type菌株数量
Strain number来源
SourceST型1)
ST type菌株数量
Strain number来源
SourceST-260 43 死胚 Dead embryo ST-3203 1 地面积水 Excess surface water ST-2100 15 死胚/孵化机
Dead embryo/IncubatorST-876 1 孵化机污垢 Incubator filth ST-3202 10 孵化机污垢 Incubator filth ST-1336 1 饲料Fodder ST-1971 5 死胚 Dead embryo ST-2640 1 死胚 Dead embryo ST-3200 2 死胚 Dead embryo ST-898 1 笼架污垢 Cage shelf filth ST-3201 2 饮水 Drinking water ST-1965 1 死胚 Dead embryo ST-3199 1 死胚 Dead embryo ST-3204 1 地面积水 Excess surface water ST-234 1 地面灰尘 Ground dust ST-1054 1 机械部件污垢
Mechanical component filthST-2552 1 笼架污垢 Cage shelf filth ST-2670 1 地面灰尘 Ground dust 1)ST-3199、ST-3200、ST-3201、ST-3202、ST-3203、ST-3204为本研究新发现的ST型
1)ST-3199, ST-3200, ST-3201 and ST-3202, ST-3203 and ST-3204 are the ST types discovered in this research -
[1] FOLIC M M, DJORDJEVIC Z, FOLIC N, et al. Epidemiology and risk factors for healthcare-associated infections caused by Pseudomonas aeruginosa[J]. Journal of Chemotherapy, 2020, 33(5): 294-301.
[2] GUPTA K K, DEVI D. Characteristics investigation on biofilm formation and biodegradation activities of Pseudomonas aeruginosa strain ISJ14 colonizing low density polyethylene (LDPE) surface[J]. Heliyon, 2020, 6(7): e04398. doi: 10.1016/j.heliyon.2020.e04398
[3] 孙理云, 孔瑞娜, 温广辉, 等. 健康肉鸡高比率携有多重耐药铜绿假单胞菌[J]. 中国人兽共患病学报, 2012, 28(2): 193-195. doi: 10.3969/j.issn.1002-2694.2012.02.025 [4] 李玲. 鸡铜绿假单胞菌的分离鉴定及生物学特性研究[D]. 保定: 河北农业大学, 2013. [5] HASSAN W H, IBRAHIM A M K, SHANY S A S, et al. Virulence and resistance determinants in Pseudomonas aeruginosa isolated from pericarditis in diseased broiler chickens in Egypt[J]. Journal of advanced veterinary and animal research, 2020, 7(3): 452-463. doi: 10.5455/javar.2020.g441
[6] SALEEM S, BOKHARI H. Resistance profile of genetically distinct clinical Pseudomonas aeruginosa isolates from public hospitals in central Pakistan[J]. Journal of Infection and Public Health, 2020, 13(4): 598-605. doi: 10.1016/j.jiph.2019.08.019
[7] MAUNDERS E A, TRINIMAN R C, WESTERN J, et al. Global reprogramming of virulence and antibiotic resistance in Pseudomonas aeruginosa by a single nucleotide polymorphism in elongation factor, fusA1[J]. Journal of Biological Chemistry, 2020, 295(48): 16411-16426. doi: 10.1074/jbc.RA119.012102
[8] 卢斌, 姚燕, 陆英, 等. 某二甲医院铜绿假单胞菌标本来源、病区分布及耐药性分析[J]. 浙江医学, 2021, 43(6): 653-655. doi: 10.12056/j.issn.1006-2785.2021.43.6.2020-3073 [9] MERRADI M, KASSAH-LAOUAR A, AYACHI A, et al. Occurrence of VIM-4 metallo-beta-lactamase-producing Pseudomonas aeruginosa in an Algerian hospital[J]. Journal of Infection in Developing Countries, 2019, 13(4): 284-290. doi: 10.3855/jidc.10679
[10] JAHAN M I, RAHAMAN M M, HOSSAIN M A, et al. Occurrence of intI1-associated VIM-5 carbapenemase and co-existence of all four classes of beta-lactamase in carbapenem-resistant clinical Pseudomonas aeruginosa DMC-27b[J]. Journal of Antimicrobial Chemotherapy, 2020, 75(1): 86-91.
[11] MARTINS W M B S, NARCISO A C, CAYO R, et al. SPM-1-producing Pseudomonas aeruginosa ST277 clone recovered from microbiota of migratory birds[J]. Diagnostic Microbiology and Infectious Disease, 2018, 90(3): 221-227. doi: 10.1016/j.diagmicrobio.2017.11.003
[12] FERNANDES M R, SELLERA F P, MOURA Q, et al. Zooanthroponotic transmission of drug-resistant Pseudomonas aeruginosa, Brazil[J]. Emerging Infectious Diseases, 2018, 24(6): 1160-1162. doi: 10.3201/eid2406.180335
[13] TREEPONG P, KOS V N, GUYEUX C, et al. Global emergence of the widespread Pseudomonas aeruginosa ST235 clone[J]. Clinical Microbiology and Infection, 2018, 24(3): 258-266. doi: 10.1016/j.cmi.2017.06.018
[14] PEREZ-VAZQUEZ M, SOLA-CAMPOY P J, ZURITA A M, et al. Carbapenemase-producing Pseudomonas aeruginosa in Spain: Interregional dissemination of the high risk-clones ST175 and ST244 carrying blaVIM-2, blaVIM-1, blaIMP-8, blaVIM-20 and blaKPC-2[J]. International Journal of Antimicrobial Agents, 2020, 56(1): 106026. doi: 10.1016/j.ijantimicag.2020.106026.
[15] XIANG Y, YAN L, ZHENG X C, et al. Rapid detection of Pseudomonas aeruginosa by cross priming amplification[J]. Journal of Integrative Agriculture, 2020, 19(10): 2523-2529. doi: 10.1016/S2095-3119(20)63187-2
[16] Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing: 28th informational CLSI document: M100-S28[S]. 2018. www.clsi.org.
[17] CURRAN B, JONAS D, GRUNDMANN H, et al. Development of a multilocus sequence typing scheme for the opportunistic pathogen Pseudomonas aeruginosa[J]. Journal of Clinical Microbiology, 2004, 42(12): 5644-5649. doi: 10.1128/JCM.42.12.5644-5649.2004
[18] DAVIS T J, KARANJIA A V, BHEBHE C N, et al. Pseudomonas aeruginosa volatilome characteristics and adaptations in chronic cystic fibrosis lung infections[J]. mSphere, 2020, 5(5): e00843-20.
[19] DAME J A, BEYLIS N, NUTTALL J, et al. Pseudomonas aeruginosa bloodstream infection at a tertiary referral hospital for children[J]. BMC Infectious Diseases, 2020, 20(1): 729. doi: 10.1186/s12879-020-05437-1.
[20] RAMOS M S, FURLAN J P R, GALLO I F L, et al. High level of resistance to antimicrobials and heavy metals in multidrug-resistant Pseudomonas sp. isolated from water sources[J]. Current Microbiology, 2020, 77(10): 2694-2701. doi: 10.1007/s00284-020-02052-w
[21] JACKSON L, DEPAS W, MORRIS A J, et al. Visualization of Pseudomonas aeruginosa within the sputum of cystic fibrosis patients[J]. Jove-Journal of Visualized Experiments, 2020(161): e61631.
[22] 秦柯君, 曹献芹, 陈派强, 等. 铜绿假单胞菌临床分布及耐药机制研究[J]. 中国病原生物学杂志, 2021, 16(2): 224-227. [23] 杨婧, 陈丽华. 铜绿假单胞菌生物被膜与宿主免疫的关系[J]. 中国微生态学杂志, 2017, 29(7): 861-865. [24] 吴振安, 张亮. 铜绿假单胞菌的临床分布及耐药性分析[J]. 中国临床医生杂志, 2021, 49(1): 55-57. doi: 10.3969/j.issn.2095-8552.2021.01.017 [25] 郑百慧, 龚春, 梅黎, 等. 呼吸与危重症医学病房分离的碳青霉烯耐药铜绿假单胞菌的分子流行病学[J]. 中华医院感染学杂志, 2020, 30(17): 2610-2614. [26] MOLINA-MORA J A, CHINCHILLA-MONTERO D, CHAVARRIA-AZOFEIFA M, et al. Transcriptomic determinants of the response of ST-111 Pseudomonas aeruginosa AG1 to ciprofloxacin identified by a top-down systems biology approach[J]. Scientific Reports, 2020, 10(1): 13717. doi: 10.1038/s41598-020-70581-2.
[27] MULET X, FERNANDEZ-ESGUEVA M, NORTE C, et al. Validation of MALDI-TOF for the early detection of the ST175 high-risk clone of Pseudomonas aeruginosa in clinical isolates belonging to a Spanish nationwide multicenter study[J]. Enfermedades Infecciosas y Microbiología Clínica, 2021, 39(6): 279-282.
[28] BOTELHO J, GROSSO F, PEIXE L. Unravelling the genome of a Pseudomonas aeruginosa isolate belonging to the high-risk clone ST235 reveals an integrative conjugative element housing a bla(GES-6) carbapenemase[J]. Journal of Antimicrobial Chemotherapy, 2018, 73(1): 77-83. doi: 10.1093/jac/dkx337


 下载:
下载: