Increase of knock-in efficiency in HEK293T cells by fusing RAD18 factor to Cas9
-
摘要:目的
在哺乳动物细胞中,RAD51和RAD18是调节同源定向修复和非同源末端连接的关键因子。本研究目的在于探讨这2个因子在eSpCas9系统中对HEK293T细胞的定点敲入(Knock-in, KI)效率的影响,进而提高KI效率。
方法通过构建eSpCas9-RAD51、eSpCas9-RAD18融合蛋白以及过表达RAD51、RAD18载体,比较它们的KI效率差异。
结果eSpCas9-RAD18系统可以显著提高HEK293T细胞KI效率,约为原来eSpCas9系统的1.4~1.9倍(P<0.01),而eSpCas9-RAD51系统和过表达RAD51/RAD18对KI效率无显著提高作用。
结论本研究构建的eSpCas9-RAD18系统可以有效地提高HEK293T细胞的KI效率,可为基因编辑、基因治疗及定点转基因提供一种新的辅助整合工具。
-
关键词:
- 基因编辑 /
- CRISPR/Cas9系统 /
- 基因敲入 /
- RAD18 /
- 融合蛋白
Abstract:ObjectiveIn mammalian cells, RAD51 and RAD18 are the key factors for regulating the relationship between non-homologous end joining and homology-directed repair. The purpose of this study was to explore the effects of these two factors on the knock-in (KI) efficiency in HEK293T cell lines by eSpCas9 system, and improve the KI efficiency.
MethodeSpCas9-RAD51, eSpCas9-RAD18 fusion proteins and RAD51, RAD18 overexpression vectors were constructed to compare their difference of KI efficiency.
ResultOnly the eSpCas9-RAD18 system could significantly increase the KI efficiency of HEK293T cells, which was about 1.4~1.9 times that of the original eSpCas9 system (P<0.01). The eSpCas9-RAD51 system and overexpression ofRAD51/RAD18 did not improve the KI efficiency.
ConclusionThe eSpCas9-RAD18 system constructed in this study can effectively improve the KI efficiency in HEK293T cells, and provides a novel auxiliary integration tool for gene editing, gene therapy and site-specific transgenesis.
-
Keywords:
- gene editing /
- CRISPR/Cas9 system /
- gene knock-in /
- RAD18 /
- fusion protein
-
草地贪夜蛾Spodoptera frugiperda是一种原产于美洲的热带和亚热带地区的杂食性害虫,严重威胁农业生产。其寄主范围广,能够为害玉米、水稻、棉花、大豆等80多种作物[1]。不同的草地贪夜蛾群体通常对特定作物具有取食趋性[2]。田间草地贪夜蛾主要分为玉米型(Corn strain)和水稻型(Rice strain)2个亚群[3]。玉米型主要取食玉米、高粱、棉花,而水稻型的取食性相对较为广谱,主要取食水稻、苜蓿、牧草、小米等作物[4]。除了取食偏好外,2个品系的草地贪夜蛾在生理特征、交配行为和性外激素的组成方面有很大的不同。但是,由于2种寄主型形态学和行为方面的相似性,在田间难以被区分开[5]。因此,对2种寄主型的草地贪夜蛾开展分子鉴定,将有助于了解该入侵物种的遗传结构,掌握害虫的入侵和扩散规律以及制定有效的防控策略[6-7]。
草地贪夜蛾亚群的分类主要根据田间采集的幼虫寄主范围,并结合分子标记的分化进行界定。基于线粒体细胞色素C氧化酶亚基I基因(COI)的分子标记不仅能够区分玉米型和水稻型2个亚群的,而且具备鉴别其亚型的能力,是目前最常用的有效分子条形码[7-10]。另外,有研究表明,根据Z染色体上编码磷酸丙糖异构酶基因(Tpi)开发的分子标记,能够更好地鉴定到草地贪夜蛾雌性“水稻型”和雄性“玉米型”的杂交型[7, 11]。2种标记有机结合能够提高鉴定的准确率[12-13]。
2018年底草地贪夜蛾通过中缅边境进入我国境内,目前已在云南、湖南、广西、广东等地发现其对玉米的为害[14]。广东省是水稻主要种植区,严密监测草地贪夜蛾是否存在为害水稻的风险显得格外重要[15]。另外,广东省为热带和亚热带季风气候区,草地贪夜蛾可在该省越冬,是可阻止其北迁的桥头堡[16]。本研究通过基于COI和Tpi 2个基因片段的分子标记,比较它们对入侵广州的草地贪夜蛾的鉴别能力,为监测虫源扩散规律以及预测预警虫情提供参考。
1. 材料与方法
1.1 供试昆虫采集
供试草地贪夜蛾于2019年5月12日在广州花都花山镇儒林村玉米地采集。寄主为喇叭口期的春玉米,所有样本均为4~6龄幼虫,6个样本分别标记为1#、2#、3#、4#、5#和6#。单头活体幼虫采集后放入含有新鲜玉米叶片的50 mL离心管,带回实验室进行形态学鉴定后,用液氮速冻储存于−80 ℃用于DNA提取。
1.2 DNA提取和扩增
采用DNA/RNA共提取试剂盒(TIANGEN,中国)分别提取单头幼虫DNA和RNA。虫体放入含有1.5 mL DNA抽提液的2 mL离心管内,加入钢珠,放入样品研磨机(Bullet Blender®,美国)处理后,按照试剂盒说明书操作。DNA使用NanoDrop分光光度计(2000,赛默飞世尔)测定质量浓度,稀释到10 ng/μL后,用作PCR扩增模板。
用于PCR扩增COI和Tpi部分基因片段的引物如表1所示,选用的PCR引物在COI基因的相对位置如图1所示。采用PrimeSTAR® Max DNA Polymerase PCR(TaKaRa,日本)进行PCR扩增,反应体系包括PrimeSTAR® Max DNA Polymerase PCR(2×)10 μL,上游引物和下游引物各0.4 μL,模板1 μL,加ddH2O至20 μL;PCR反应为:98 ℃预变性2 min;98 ℃变性10 s,60 ℃退火15 s,72 ℃延伸30 s,35个循环;72 ℃延伸5 min。
表 1 扩增COI和Tpi基因片段所用引物Table 1. Primers used for amplifying COI and Tpi gene fragments使用15 g/L的琼脂糖凝胶电泳检测扩增片段特异性后,PCR产物由广州天一辉远基因科技有限公司进行切胶测序。对于测序结果为双峰或者低质量的样品,则将其PCR产物克隆到pEASY®-Blunt Cloning Vector后,每个样品各挑取10个克隆进行测序。
1.3 序列分析
利用Lastergene软件包对正反向测序序列进行人工拼接、峰型校正和序列比对。从GenBank数据库下载已经发布的草地贪夜蛾COI和Tpi基因序列。针对已报道的基于COI和Tpi基因序列的“水稻型”和“玉米型”单倍型特点,分别进行差异位点分析,确定本研究中草地贪夜蛾的具体亚型。
2. 结果与分析
2.1 COI标记鉴定草地贪夜蛾单倍型
利用标记COI-A,COI-B和COI-C对所采集的6个样品进行了寄主型分析。结果(图2)显示,有5个样品扩增出了预期大小的单一特异性条带,将其PCR产物用于直接测序。
对COI-A的序列分析表明,来自4个样品(1#、2#、4#、6#)的序列与已报道的草地贪夜蛾水稻型(HM136593)仅存在一处碱基差异,与玉米型(HM136586)存在8处单碱基差异,说明根据COI-A标记鉴定的寄主型与实际样品来源寄主型并不一致(图3);对样品3#的COI-A产物测序分析发现其为双峰杂合,克隆重新测序后,分别获得了样品3#-1和3#-2两种单倍型。其中,3#-1与上述4个样品检测到的单倍型完全一致,3#-2则与Mythimna separate(KX863042) 对应的片段序列上存在1个碱基的差异,说明3#样品可能是草地贪夜蛾和M. separate的杂合(图3)。
进一步用COI-B和COI-C标记对样品进行寄主型鉴定,结果表明,除了样品5#无法扩增出条带之外,其余的5个样品均扩增出单一的预期条带。PCR产物测序结果显示所测样品序列完全一致且为水稻型(图2)。
将COI-A、COI-B和COI-C等3个标记的结果相比较,发现COI-A能够区分出草地贪夜蛾和M. separate的2种单倍型,而COI-B和COI-C仅能区分出草地贪夜蛾1种单倍型,说明虽然基于COI基因片段序列鉴定到的草地贪夜蛾寄主型与实际采集寄主种类不相符,但是,COI-A标记具有较好的物种鉴别能力,能够区分出杂合类型。
2.2 TPi标记鉴定草地贪夜蛾单倍型
基于Tpi基因片段对所采集的6个样品进行了寄主型分析。除了样品5#无法扩增出条带之外,其余的5个样品均扩增出了预期大小的单一特异性条带(图4)。
对PCR产物进行双向测序并分析其序列差异,结果显示,所检测的序列与已报道玉米型一致,说明基于Tpi基因片段的鉴定结果与样品的寄主来源是一致的。样品2#和样品3#,在第126和127碱基处出现GA/AT的杂合双峰(图5),说明入侵广东的草地贪夜蛾种类并不单一。
3. 结论与讨论
本研究采集了入侵广州市的草地贪夜蛾样本,利用基于COI和Tpi基因片段的分子标记进行了寄主型的鉴定。基于COI-A、COI-B和COI-C标记的鉴定结果均显示入侵广州的草地贪夜蛾寄主单倍型为水稻型,与实际采集寄主不相符,可能原因是该标记不适合于检测入侵广州的草地贪夜蛾群体。但基于COI-A的标记能够区分草地贪夜蛾与其他近源种的基因型,为研究入侵草地贪夜蛾的田间行为和扩散提供丰富的信息。而COI-B和COI-C只鉴定到1种单倍型。COI-A和COI-B标记事实上扩增的是COI基因的同一片段,差别在于COI-A标记扩增的片段较小,且使用的是简并引物。因此,COI-A标记具有比COI-B标记和COI-C标记更强的鉴别能力。
Tpi基因标记的鉴定结果显示入侵广州的草地贪夜蛾寄主型为玉米型,与实际采集的寄主一致。基于Tpi基因标记,入侵广州的草地贪夜蛾群体被检测到具有2种单倍亚型,在该片段第126和第127位碱基具有RW杂合。先前报道的入侵云南的群体只有一种单倍型[12],说明草地贪夜蛾可能在扩散的过程中出现遗传杂交现象。
草地贪夜蛾是联合国粮农组织全球预警的跨国界迁飞性重大害虫。目前,入侵我国的草地贪夜蛾主要为害玉米,未见在水稻上定殖为害。华南地区是我国水稻主产区,应慎重评估草地贪夜蛾为害水稻的风险。对于“玉米型”和“水稻型”的区分,仅依靠个别基因的片段进行评估是不全面的[6]。基于微卫星标记的分子指纹分析可以较全面地了解草地贪夜蛾的遗传结构动态变化规律[17]。获得与寄主取食相关基因的信息进而开发基因功能性分子标记,是精准鉴定草地贪夜蛾寄主单倍型的前提[18]。
-
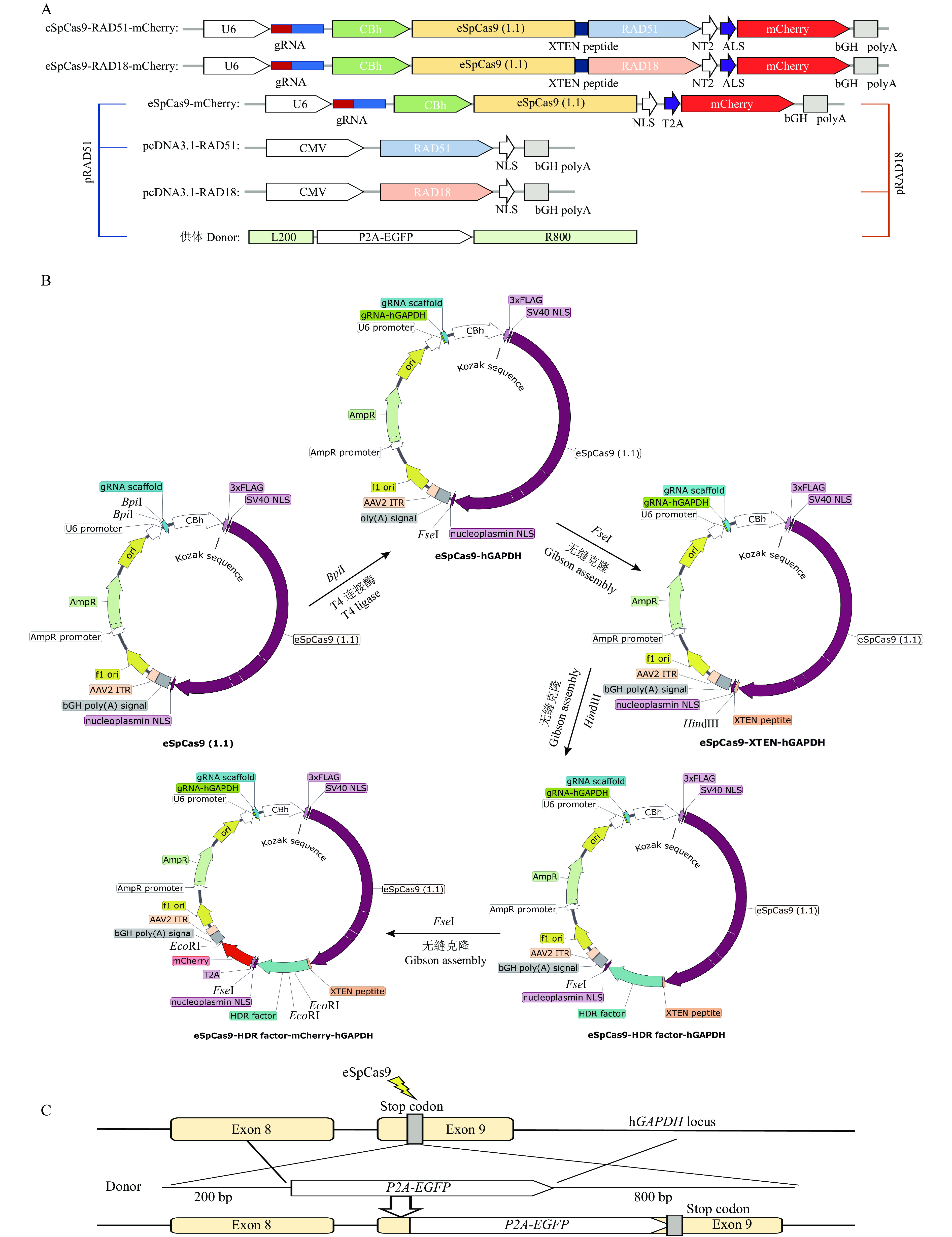
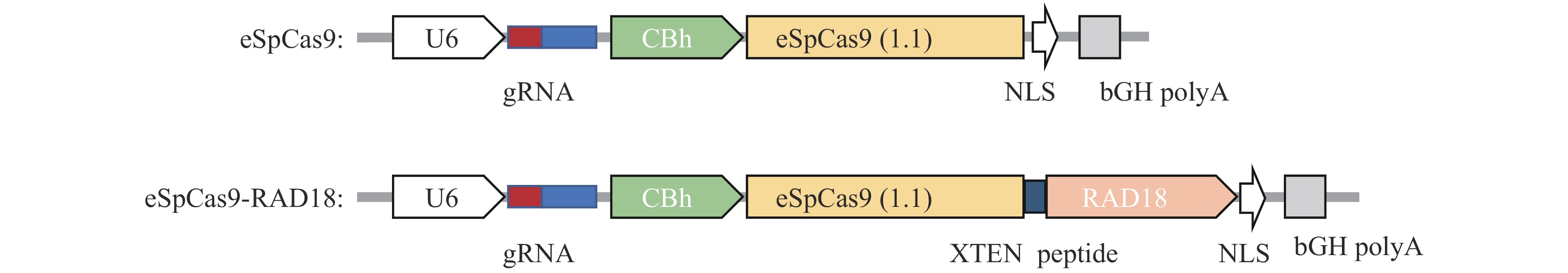
图 1 不同eSpCas9系统的构建及KI模式图
A:eSpCas9-HDR-mCherry载体和pcDNA3.1-HDR过表达载体模式图,其中U6、CBh和CMV为启动子, NLS为核定位信号, T2A和XTEN peptide为连接肽,bGH polyA为终止子, L200为靶位点处200 bp的左同源臂,R800为靶位点处800 bp的右同源臂,mCherry用于矫正eSpCas9的转染效率;B:eSpCas9相关载体构建流程;C:目的基因(P2A-EGFP)整合在人GAPDH位点的模式图,其中eSpCas9能够特异性切割基因组
Figure 1. Construction of different eSpCas9 systems and KI pattern diagram
A: Pattern diagram of eSpCas9-HDR-mCherry vector and pcDNA3.1-HDR overexpression vector, where U6,CBh and CMV are promoters, NLS is the nuclear localization sequence, T2A and XTEN peptide are linking peptides, bGH polyA is the terminator, L200 is the 200 bp left homologous arm of the target site, R800 is the 800 bp right homologous arm of the target site, and mCherry is used to correct the transfection efficiency of eSpCas9; B:eSpCas9 related vector construction process; C: A schematic diagram of the target gene (P2A-EGFP) integrated into the human GAPDH site, where eSpCas9 can specifically cleave the genome
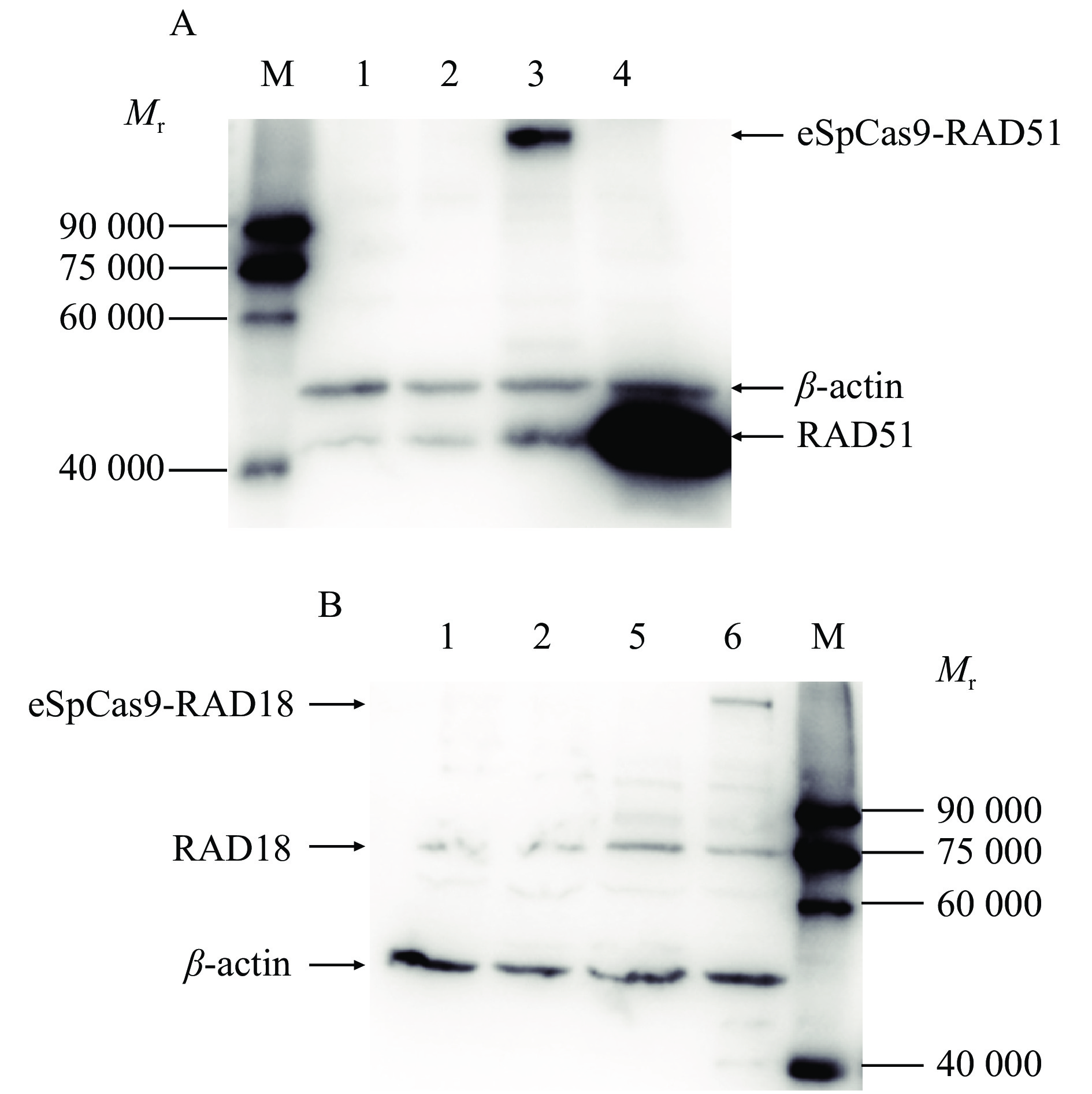
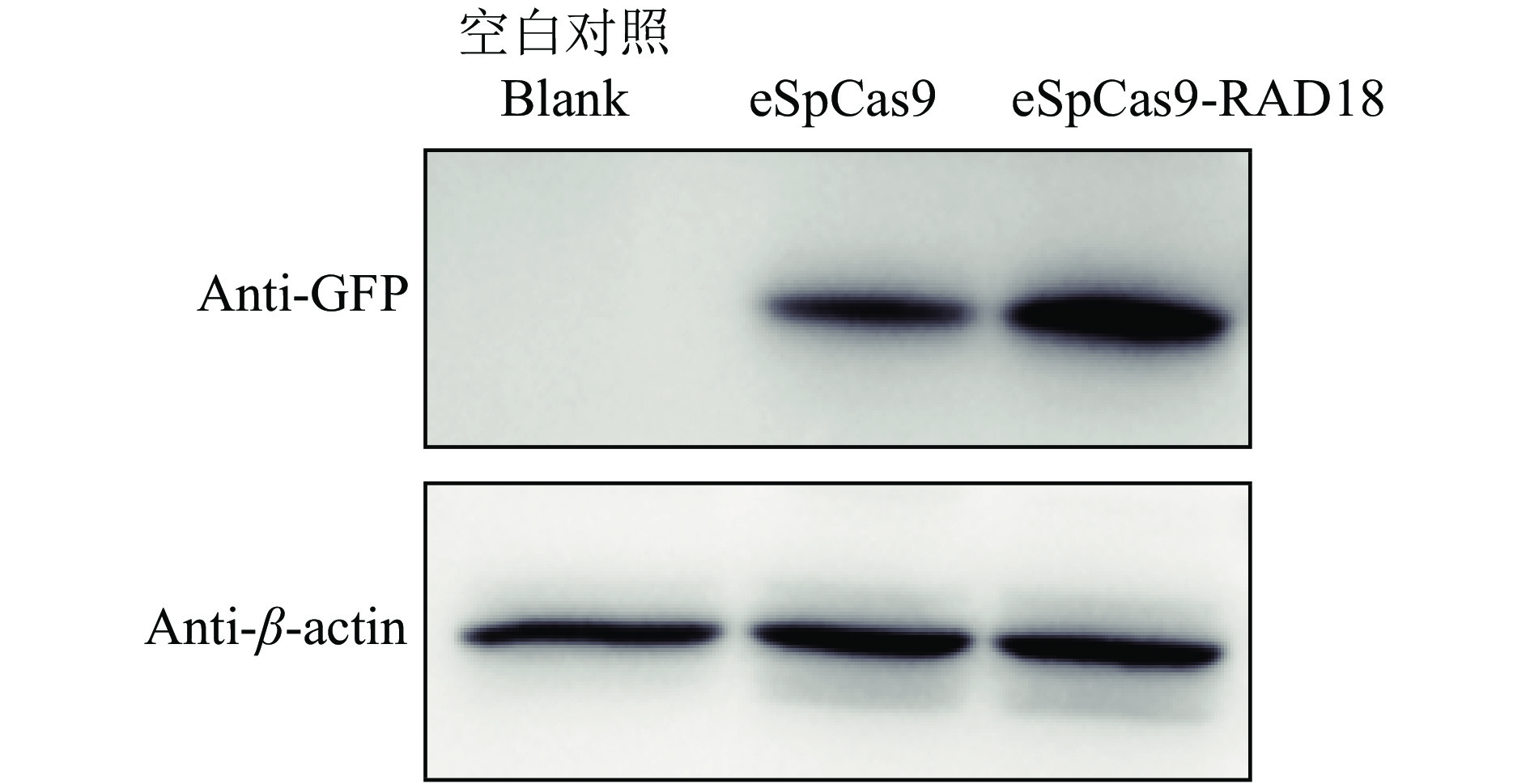
图 3 不同融合蛋白的表达结果
M:蛋白质相对分子质量标准,1:空白对照,2:eSpCas9, 3:eSpCas9-RAD51,4:pRAD51,5:pRAD18,6:eSpCas9-RAD18;pRAD51/ pRAD18 中pcDNA3.1-RAD51/RAD18的质粒用量为250 ng
Figure 3. The expression results of different fusion proteins
M: Protein marker,1: Blank,2: eSpCas9, 3: eSpCas9-RAD51,4: pRAD51,5: pRAD18,6: eSpCas9-RAD18;In pRAD51/ pRAD18, the amount of pcDNA3.1-RAD51/RAD18 plasmid is 250 ng
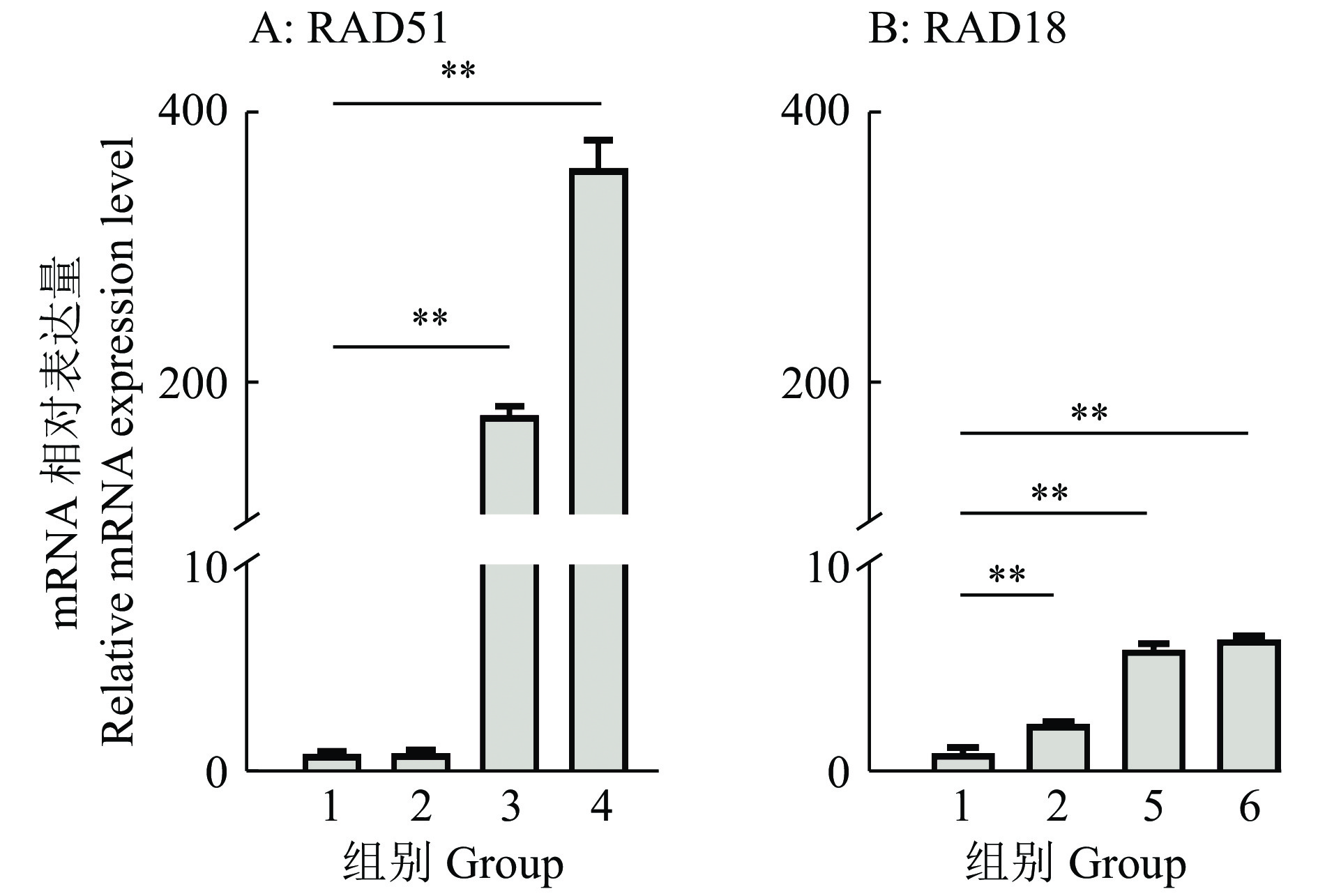
图 4 RAD51/18过表达后mRNA表达水平
1:空白对照,2:eSpCas9,3和4分别为使用100和250 ng pcDNA3.1-RAD51质粒的pRAD51组别,5和6分别为使用100和250 ng pcDNA3.1-RAD18质粒的pRAD18组别;“**”表示差异显著(P<0.01, t检验)
Figure 4. mRNA expression level after RAD51/18 overexpression
1: Blank,2: eSpCas9,3 and 4 are pRAD51 groups with 100 and 250 ng pcDNA3.1-RAD51 plasmid respectively, 5 and 6 are pRAD18 groups with 100 and 250 ng pcDNA3.1-RAD18 plasmid respectively; "**" indicates significant difference (P<0.01, t test)
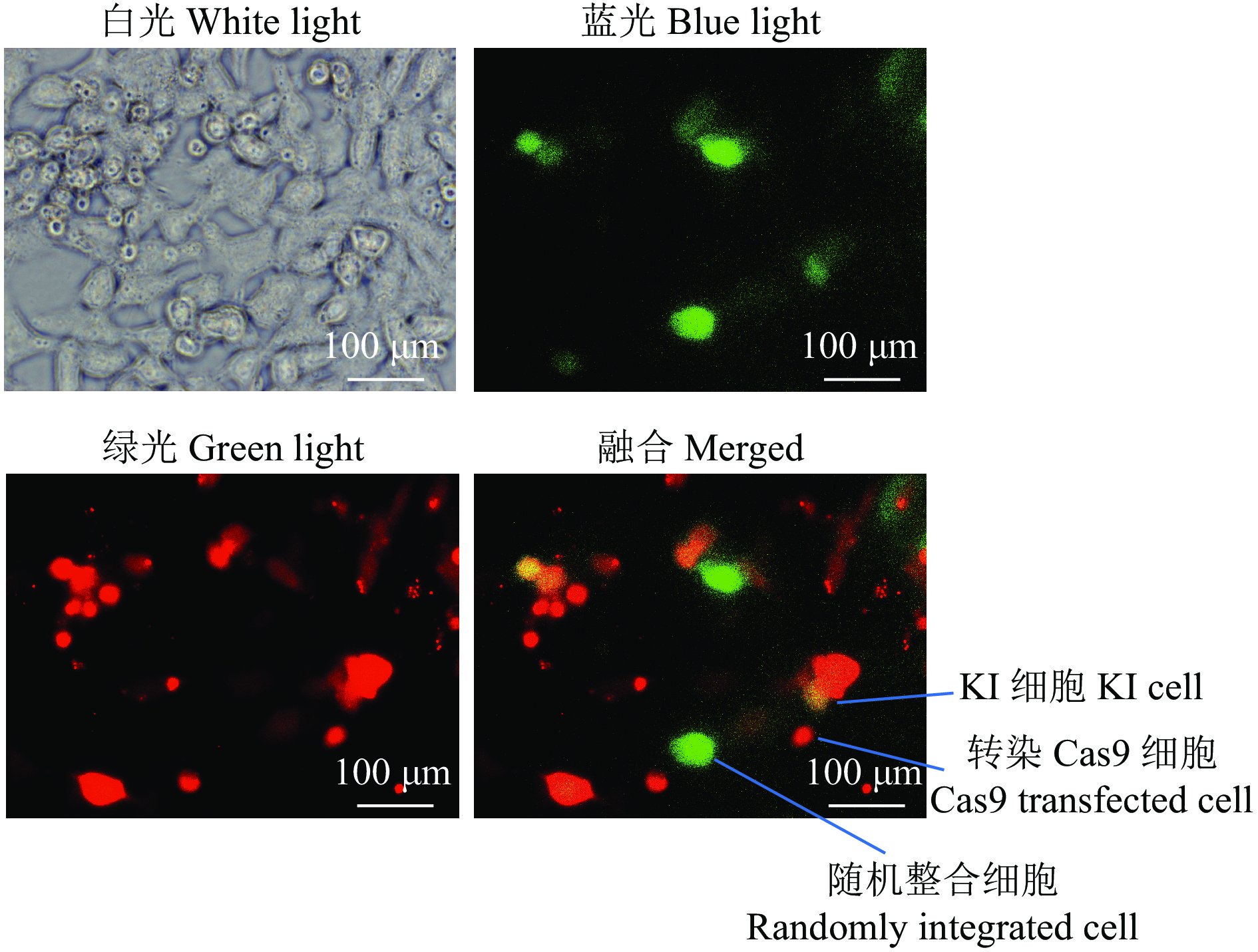
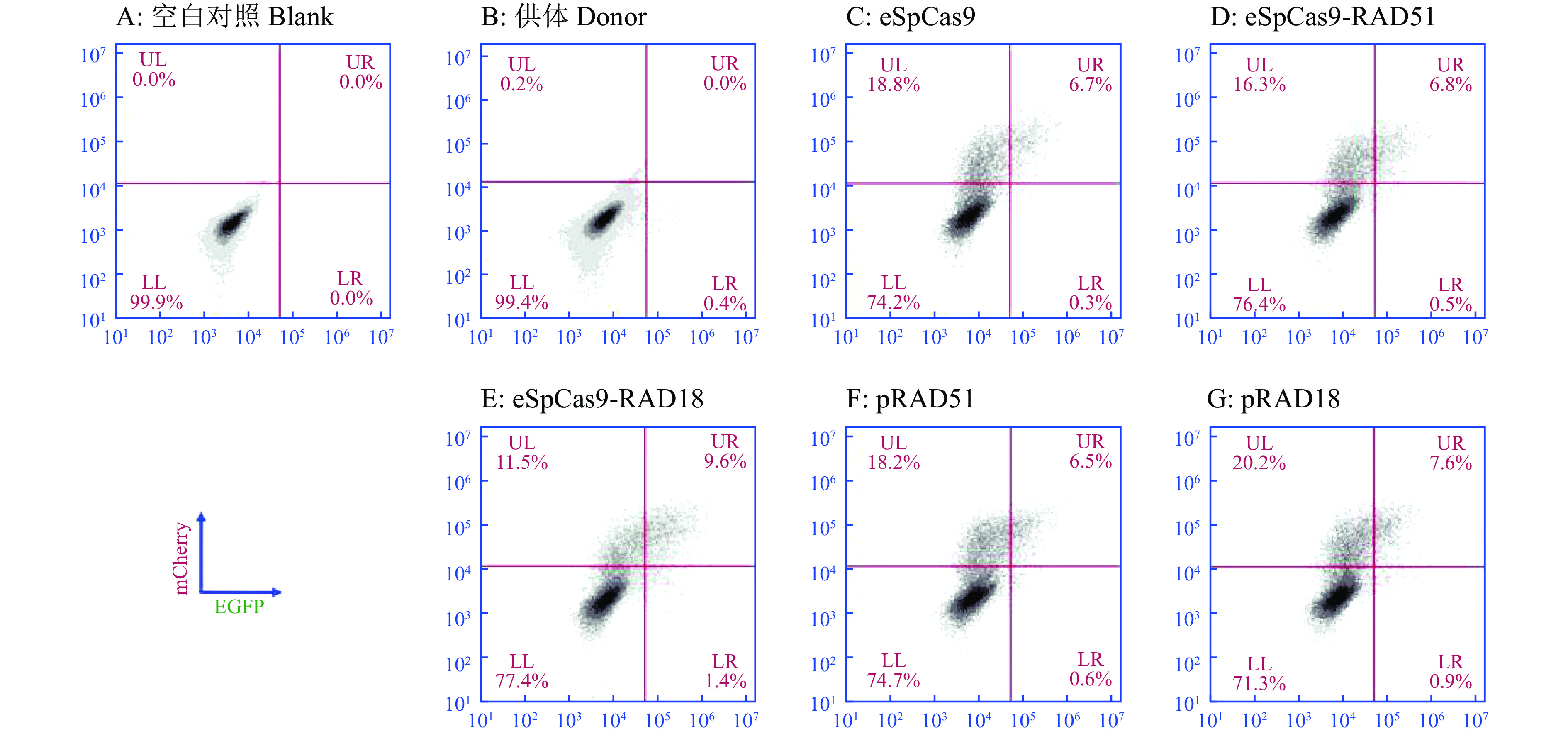
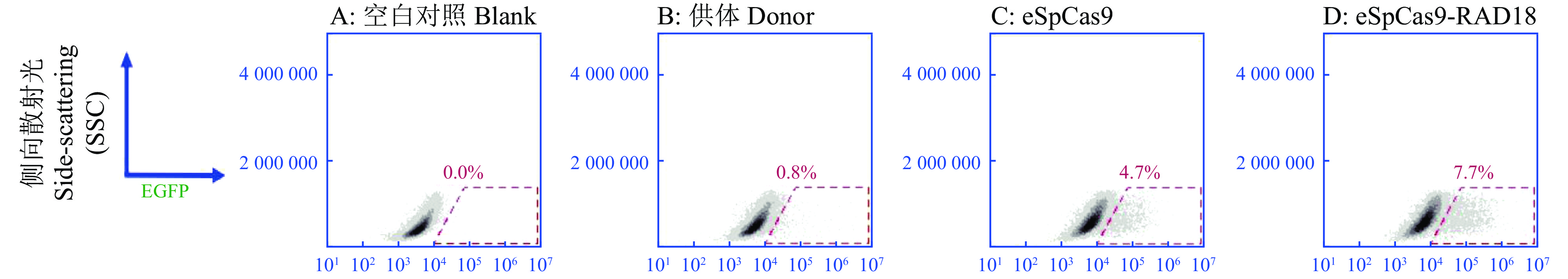
图 6 eSpCas9-HDR-mCherry系统对基因KI效率的影响
UL是仅表达红色荧光的细胞占比,UR是既表达红色荧光又表达绿色荧光的细胞占比,LL是不表达任何荧光的细胞占比,LR是仅表达绿色荧光的细胞占比
Figure 6. The effect of eSpCas9-HDR-mCherry system on gene KI efficiency
UL means the proportion of cells that only expresses red fluorescence, UR means the proportion of that expresses both red and green fluorescence , LL means the proportion of cells that do not express fluorescence, and LR means the proportion of cells that only expresses green fluorescence
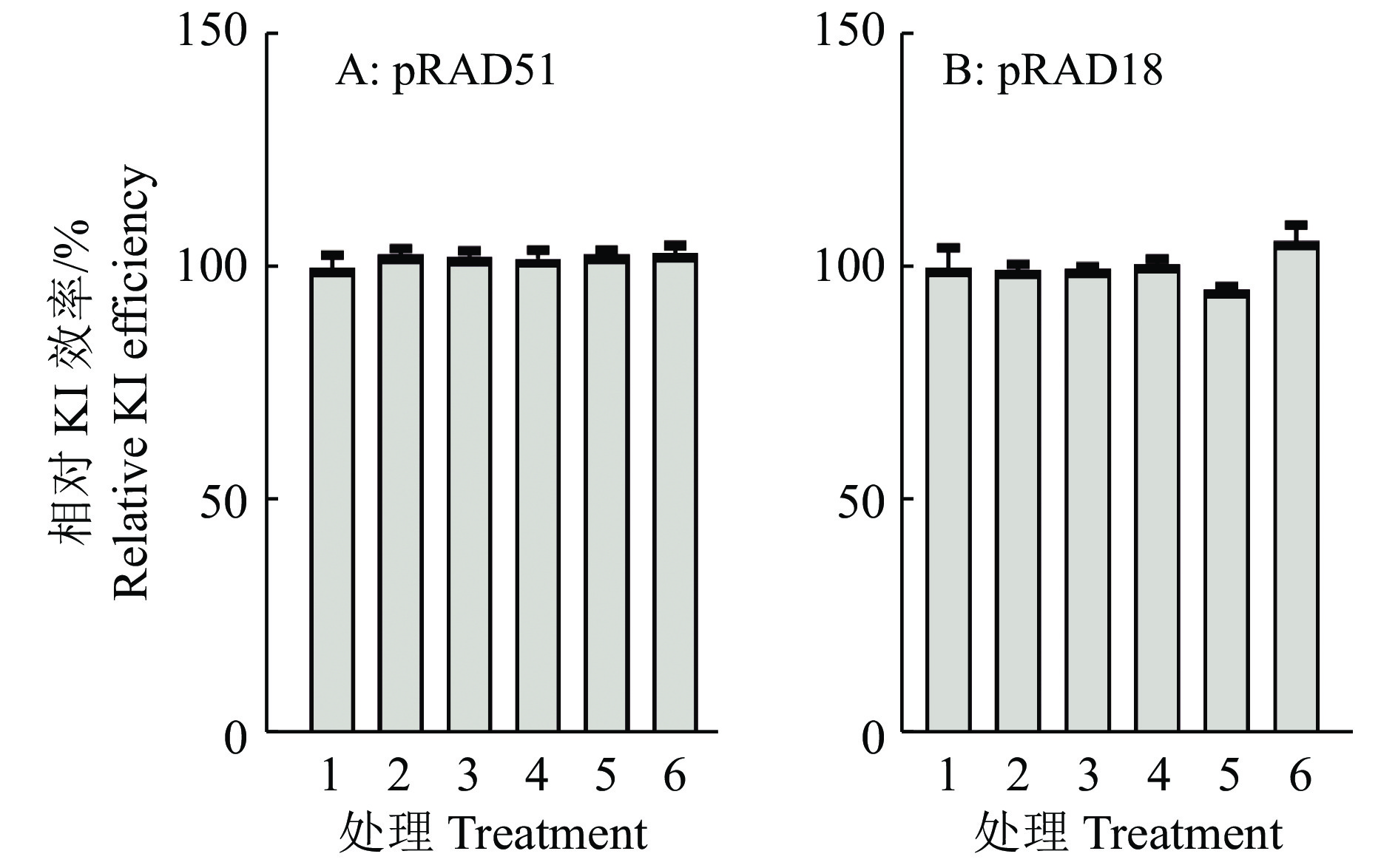
图 7 过表达不同浓度HDR因子对KI效率的影响
相对KI效率=该组别的KI效率/Control的KI效率×100%;1~6表示pRAD51/pRAD18组别中pcDNA3.1-RAD51/ pcDNA3.1-RAD18质粒的用量分别为0、17、33、100、250和500 ng
Figure 7. The effect of overexpression of HDR factors with different concentrations on the KI efficiency
Relative KI efficiency=KI efficiency of the group/KI efficiency of control×100%;1−6 indicate the amounts of pcDNA3.1-RAD51/ pcDNA3.1-RAD18 plasmid in pRAD51/pRAD18 group are 0,17,33,100,250 and 500 ng
表 1 引物序列和用途
Table 1 Sequence and use of primer
引物名称
Primer name正向序列 (5′→3′)
Forward sequence反向序列 (5′→3′)
Reverse sequence用途
UsegRNA-hGAPDH CACCAGCCCCAGCAAGAGCACAAG AAACCTTGTGCTCTTGCTGGGGCT gRNA载体构建
gRNA vector constructionL200R800 AACGACCACTTTGTCAAGCTCAT ACCCTGACACATCTCAGTTCATCTT 转染供体
Transfection donorXTEN CAGCTGGGAGGCGACAGCGGTTCAGAGACCCCAGGAACTAGCGAGAGCGCTACACCGGAATCG TTCTTTTTTGCCTGGCCGGCCTTTTTCGTGGCCGCCGGCCTTTTAAGCTTCGATTCCGGTGTAGCGCTCT Cas9过度载体构建
Cas9 excessive vector constructionIF-Cas9-RAD51 GCTACACCGGAATCGATGGCAATGCAGATGCAGCT GGCCGCCGGCCTTTTGTCTTTGGCATCTCCCACTCCAT Cas9-HDR载体构建Cas9-HDR vector construction IF-Cas9-RAD18 GCTACACCGGAATCGATGGACTCCCTGGCCGAGTC GGCCGCCGGCCTTTTATTCCTATTACGCTTGTTTCTTGGTTC IF-RAD51-NLS TTGGTACCGAGCTCGGATCCGCCACCATGGCAATGCAGATGCAGCT CCTCTAGACTCGAGCGGCCGCTTACTTTTTCTTTTTTGCCTGGCCGG 过表达载体构建
Overexpression vector constructionIF-RAD18-NLS TTGGTACCGAGCTCGGATCCGCCACCATGGACTCCCTGGCCGAGTC CCTCTAGACTCGAGCGGCCGCTTACTTTTTCTTTTTTGCCTGGCCGG IF-T2A-mCherry CGGCCACGAAAAAGGCCGGCCAGGCAAAAAAGAAAAAGCTTGAGG CTCTAGGAATTCTTACTTGTACAGCTCGTCCATGCC hU6 CCGTAACTTGAAAGTATTTCG Sanger测序
Sanger sequencingBGH TAGAAGGCACAGTCGAGG T7 TAATACGACTCACTATAGGG mCherry TTGGTCACCTTCAGCTTGG JD-KI-hGAPDH ACCACAGTCCATGCCATCACT AGTCGTGCTGCTTCATGTGGT KI细胞鉴定
KI cell identificationq-ACTB CCAACCGCGAGAAGATGACC AGTCCATCACGATGCCAGT 荧光定量PCR
Fluorescence quantitative PCRq-CtIP ACAGGAACGAATCTTAGATGCACA ACATAAGCCTGCTCTTAACCGAT q-RAD50 AGATCGCCATCAAGAACATATCCG TCACTGAATGGTCCACGCTCA q-RAD51 CCTGCCAGCTTCCCATTGACC CGTTCTGGCCTAAAGGTACCCTC q-KU70 TGCCAACCTCTTTAGTGATGTCC CCCAGGTTTCTTCAGGTGCAT q-LIG4 AACCATACAGCAAGTAAACGACCT TCCGTATAAGCCACTTTTGCTCA q-EGFP ACCACTACCAGCAGAACACC CATGTGATCGCGCTTCTCGTT 表 2 mCherry蛋白矫正不同eSpCas9系统的KI效率1)
Table 2 The effect of different eSpCas9 systems on KI efficiency
组别
GroupCas9载体转染效率/%
Transfection efficiency of Cas9 vector
(UL+UR)KI效率/% KI efficiency 原始值
Original (UR)校正值
Corrected [UR/(UL+UR)]空白对照 Blank — 0 — 供体 Donor — 0.35±0.03 — eSpCas9 24.25±1.33 6.71±0.33 27.69±1.00 eSpCas9-RAD51 22.87±0.60 6.63±0.11 29.01±1.02 eSpCas9-RAD18 20.36±0.99** 9.35±0.50** 45.91±0.51** pRAD51 25.20±0.71 6.73±0.50 26.69±1.23 pRAD18 26.68±1.08* 7.08±0.95 26.43±2.69 1) UL是仅表达红色荧光的细胞占比,UR是既表达红色荧光又表达绿色荧光的细胞占比,即原始效率,LL是不表达任何荧光的细胞占比,LR是仅表达绿色荧光的细胞占比,UL+UR代表eSpCas9转进去的细胞占比,UR/(UL+UR)是矫正后的KI效率;由于Blank和Donor未转染eSpCas9载体,故在表中用“—”表示;“*”和“**”分别表示与eSpCas9组在0.05和0.01水平差异显著(单因素ANOVA分析)
1) UL means the proportion of cells that only express red fluorescent, UR means the proportion of cells that express both red and green fluorescence cells, which is the original efficiency, LL means the proportion of cells that do not express fluorescence, LR means the proportion of cells that only express green fluorescence, UL+UR represents the proportion of cells transferred by eSpCas9, and UR/(UL+UR) refers to the corrected KI efficiency; Since Blank and Donor were not transfected with eSpCas9 vector, they are indicated by “—” in the table; “*” and “**” represent significant difference from eSpCas9 group at 0.05 and 0.01 levels respectively(ANOVA) -
[1] YAN S, TU Z, LIU Z, et al. A huntingtin knockin pig model recapitulates features of selective neurodegeneration in Huntington's disease[J]. Cell, 2018, 173(4): 989-1002.e13. doi: 10.1016/j.cell.2018.03.005
[2] 史梦然, 沈宗毅, 张楠, 等. CRISPR/Cas9系统在疾病研究和治疗中的应用[J]. 生物工程学报, 2021, 37(4): 1205-1228. [3] ZHOU H, SU J, HU X, et al. Glia-to-neuron conversion by CRISPR-CasRx alleviates symptoms of neurological disease in mice[J]. Cell, 2020, 181(3): 590-603.e16. doi: 10.1016/j.cell.2020.03.024
[4] ZHANG X, LI Z, YANG H, et al. Novel transgenic pigs with enhanced growth and reduced environmental impact[J]. Elife, 2018, 7: e34286. doi: 10.7554/eLife.34286
[5] 钟翠丽. 制备一种共表达多个酶基因的环保型转基因猪[D]. 广州: 华南农业大学, 2018. [6] 严振鑫, 徐冬一. DNA双链断裂的非同源末端连接修复[J]. 生命科学, 2014, 26(11): 1157-1165. [7] LIEBER M R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway[J]. Annual Review of Biochemistry, 2010, 79: 181-211. doi: 10.1146/annurev.biochem.052308.093131
[8] LANS H, MARTEIJN J A, VERMEULEN W. ATP-dependent chromatin remodeling in the DNA-damage response[J]. Epigenetics and Chromatin, 2012, 5: 4. doi: 10.1186/1756-8935-5-4
[9] FUJITANI Y, YAMAMOTO K, KOBAYASHI I. Dependence of frequency of homologous recombination on the homology length[J]. Genetics, 1995, 140(2): 797-809. doi: 10.1093/genetics/140.2.797
[10] SABERI A, HOCHEGGER H, SZUTS D, et al. RAD18 and poly(ADP-ribose) polymerase independently suppress the access of nonhomologous end joining to double-strand breaks and facilitate homologous recombination-mediated repair[J]. Molecular and Cellular Biology, 2007, 27(7): 2562-2571. doi: 10.1128/MCB.01243-06
[11] BURROWS A E, ELLEDGE S J. How ATR turns on: TopBP1 goes on ATRIP with ATR[J]. Genes & Development, 2008, 22(11): 1416-1421.
[12] BARKLEY L R, PALLE K, DURANDO M, et al. c-Jun N-terminal kinase-mediated Rad18 phosphorylation facilitates Polη recruitment to stalled replication forks[J]. Molecular Biology of The Cell, 2012, 23(10): 1943-1954. doi: 10.1091/mbc.e11-10-0829
[13] BI X, BARKLEY L R, SLATER D M, et al. Rad18 regulates DNA polymerase kappa and is required for recovery from S-phase checkpoint-mediated arrest[J]. Molecular and Cellular Biology, 2006, 26(9): 3527-3540. doi: 10.1128/MCB.26.9.3527-3540.2006
[14] SASATANI M, XU Y, KAWAI H, et al. RAD18 activates the G2/M checkpoint through DNA damage signaling to maintain genome integrity after ionizing radiation exposure[J]. Plos One, 2015, 10(2): e0117845. doi: 10.1371/journal.pone.0117845
[15] INAGAKI A, VAN CAPPELLEN W A, VAN DER LAAN R, et al. Dynamic localization of human RAD18 during the cell cycle and a functional connection with DNA double-strand break repair[J]. DNA Repair, 2009, 8(2): 190-201. doi: 10.1016/j.dnarep.2008.10.008
[16] LIU T, HUANG J, CHEN J J. RAD18 lives a double life: Its implication in DNA double-strand break repair[J]. DNA Repair, 2010, 9(12): 1241-1248. doi: 10.1016/j.dnarep.2010.09.016
[17] SONG J, YANG D, XU J, et al. RS-1 enhances CRISPR/Cas9- and TALEN-mediated knock-in efficiency[J]. Nature Communications, 2016, 7: 10548. doi: 10.1038/ncomms10548
[18] HE X, TAN C, WANG F, et al. Knock-in of large reporter genes in human cells via CRISPR/Cas9-induced homology-dependent and independent DNA repair[J]. Nucleic Acids Research, 2016, 44(9): e85.
[19] LIVAK K J, SCHMITTGEN T D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method[J]. Methods, 2001, 25(4): 402-408.
[20] NAMBIAR T S, BILLON P, DIEDENHOFEN G, et al. Stimulation of CRISPR-mediated homology-directed repair by an engineered RAD18 variant[J]. Nature Biotechnology, 2019, 10(1): 3395.
[21] ZEMAN M K, LIN J R, FREIRE R, et al. DNA damage-specific deubiquitination regulates Rad18 functions to suppress mutagenesis[J]. Journal of Cell Biology, 2014, 206(2): 183-197. doi: 10.1083/jcb.201311063
[22] HELCHOWSKI C M, SKOW L F, ROBERTS K H, et al. A small ubiquitin binding domain inhibits ubiquitin-dependent protein recruitment to DNA repair foci[J]. Cell Cycle, 2013, 12(24): 3749-3758. doi: 10.4161/cc.26640
[23] CHARPENTIER M, KHEDHER A H Y, MENORET S, et al. CtIP fusion to Cas9 enhances transgene integration by homology-dependent repair[J]. Nature Communications, 2018, 9(1): 1133. doi: 10.1038/s41467-018-03475-7
[24] TRAN N T, BASHIR S, LI X, et al. Enhancement of precise gene editing by the association of Cas9 with homologous recombination factors[J]. Frontiers in Genetics, 2019, 10: 365. doi: 10.3389/fgene.2019.00365
[25] REES H A, YEH W H, LIU D R. Development of hRAD51-Cas9 nickase fusions that mediate HDR without double-stranded breaks[J]. Nature Communications, 2019, 10(1): 2212. doi: 10.1038/s41467-019-09983-4
[26] KURIHARA T, KOUYAMA-SUZUKI E, SATOGA M, et al. DNA repair protein RAD51 enhances the CRISPR/Cas9-mediated knock-in efficiency in brain neurons[J]. Biochemical and Biophysical Research Communications, 2020, 524(3): 621-628. doi: 10.1016/j.bbrc.2020.01.132
[27] TAKAYAMA K, IGAI K, HAGIHARA Y, et al. Highly efficient biallelic genome editing of human ES/iPS cells using a CRISPR/Cas9 or TALEN system[J]. Nucleic Acids Research, 2017, 45(9): 5198-5207. doi: 10.1093/nar/gkx130
[28] PAFFETT K S, CLIKEMAN J A, PALMER S, et al. Overexpression of RAD51 inhibits double-strand break-induced homologous recombination but does not affect gene conversion tract lengths[J]. DNA Repair, 2005, 4(6): 687-698. doi: 10.1016/j.dnarep.2005.03.003
[29] KIM P M, ALLEN C, WAGENER B M, et al. Overexpression of human RAD51 and RAD52 reduces double-strand break-induced homologous recombination in mammalian cells[J]. Nucleic Acids Research, 2001, 29(21): 4352-4360. doi: 10.1093/nar/29.21.4352
[30] THOMPSON L H, SCHILD D. Homologous recombinational repair of DNA ensures mammalian chromosome stability[J]. Mutation Research-fundamental and Molecular Mechanisms of Mutagenesis, 2001, 477(1/2): 131-153. doi: 10.1016/S0027-5107(01)00115-4
[31] LISBY M, BARLOW J H, BURGESS R C, et al. Choreography of the DNA damage response: Spatiotemporal relationships among checkpoint and repair proteins[J]. Cell, 2004, 118(6): 699-713. doi: 10.1016/j.cell.2004.08.015
[32] HUANG J, HUEN M S, KIM H, et al. RAD18 transmits DNA damage signalling to elicit homologous recombination repair[J]. Nature Cell Biology, 2009, 11(5): 592-603.
[33] KOBAYASHI S, KASAISHI Y, NAKADA S, et al. Rad18 and Rnf8 facilitate homologous recombination by two distinct mechanisms, promoting RAD51 focus formation and suppressing the toxic effect of nonhomologous end joining[J]. Oncogene, 2015, 34(33): 4403-4411. doi: 10.1038/onc.2014.371
[34] BRANZEI D, VANOLI F, FOIANI M. SUMOylation regulates Rad18-mediated template switch[J]. Nature, 2008, 456(7224): 915-920. doi: 10.1038/nature07587
[35] WONG R P, AGUISSA-TOURE A H, WANI A A, et al. Elevated expression of Rad18 regulates melanoma cell proliferation[J]. Pigment Cell & Melanoma Research, 2012, 25(2): 213-218.
-
期刊类型引用(4)
1. 邓文林. 乡土阔叶树种示范林建设——以韶关市国有仁化林场为例. 农村科学实验. 2024(18): 133-135 .  百度学术
百度学术
2. 何潇,雷相东,段光爽,丰庆荣,张逸如,冯林艳. 气候变化对落叶松人工林生物量生长的影响模拟. 南京林业大学学报(自然科学版). 2023(03): 120-128 .  百度学术
百度学术
3. 康梓杭,倪苗,邓晶,吴庆书. 海口市10种行道树最优生长模型研究. 热带生物学报. 2020(01): 79-83 .  百度学术
百度学术
4. 曹磊,刘晓彤,李海奎,雷渊才. 广东省常绿阔叶林生物量生长模型. 林业科学研究. 2020(05): 61-67 .  百度学术
百度学术
其他类型引用(9)


 下载:
下载: