Sensitivity of four phages to host bacterium Ralstonia solanacearum and analysis of their receptors
-
摘要:目的
明确不同噬菌体混合对茄劳尔氏菌(青枯菌)Ralstonia solanacearum裂解能力和对噬菌体抗性产生的影响,了解4种噬菌体的作用受体。
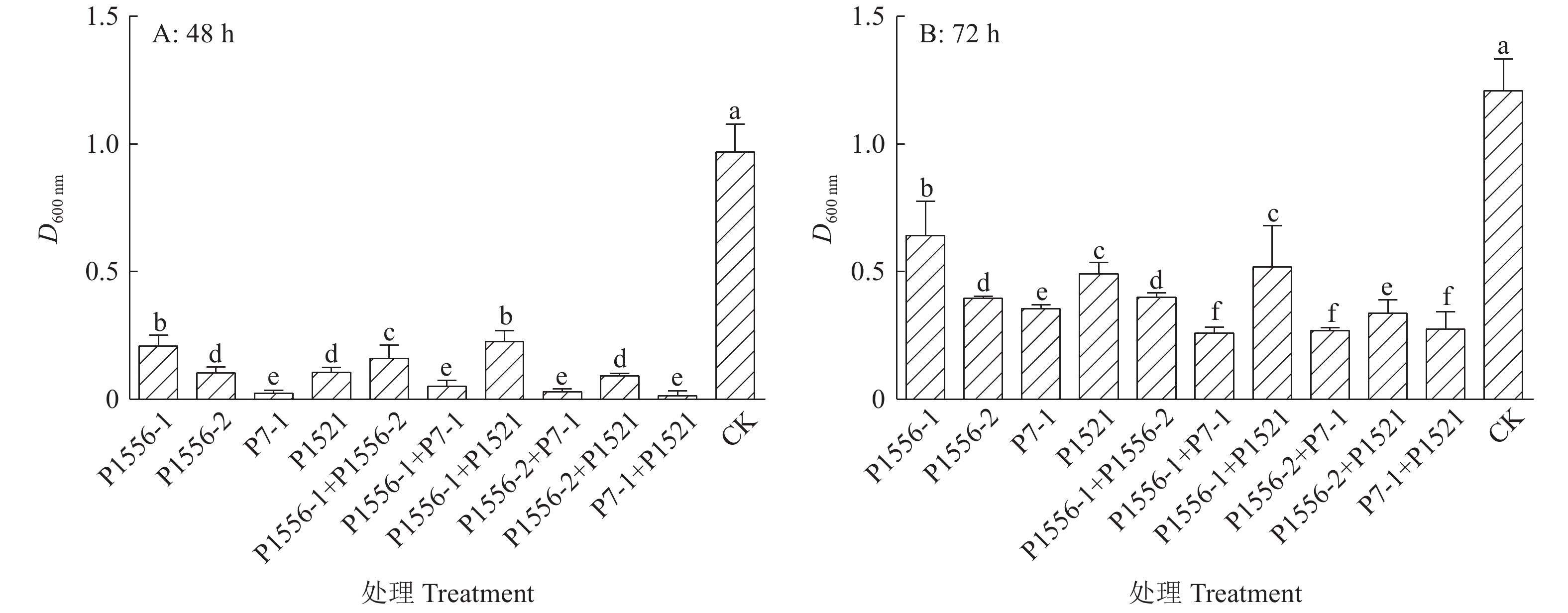
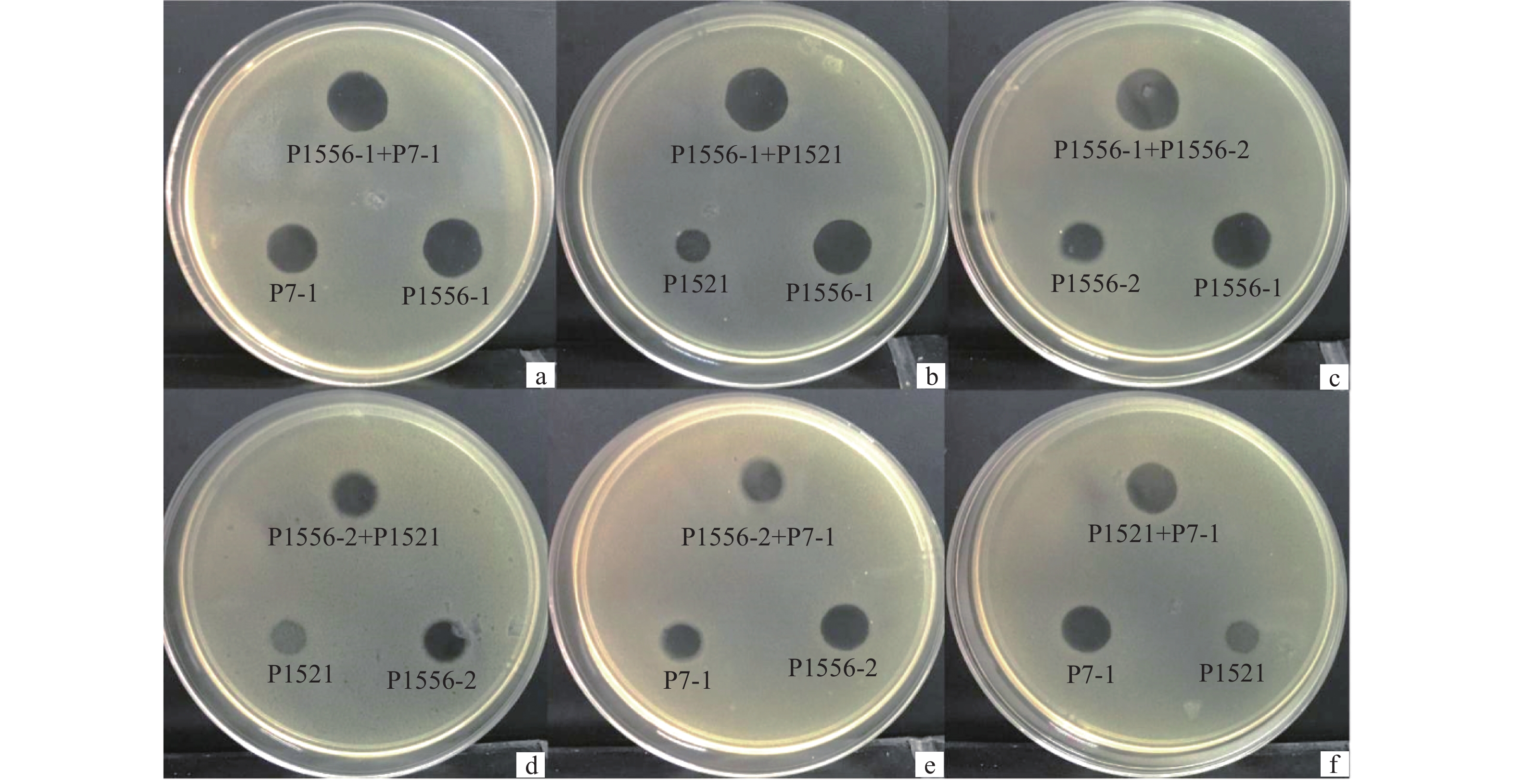
方法将4种噬菌体P1556-1、P1556-2、P7-1和P1521两两混合后,比较其在青枯菌平板上产生的噬菌斑大小;噬菌体与青枯菌混合培养测定青枯菌对噬菌体的抗性;通过噬菌体与脂多糖和膜蛋白的吸附试验测定噬菌体的作用受体。
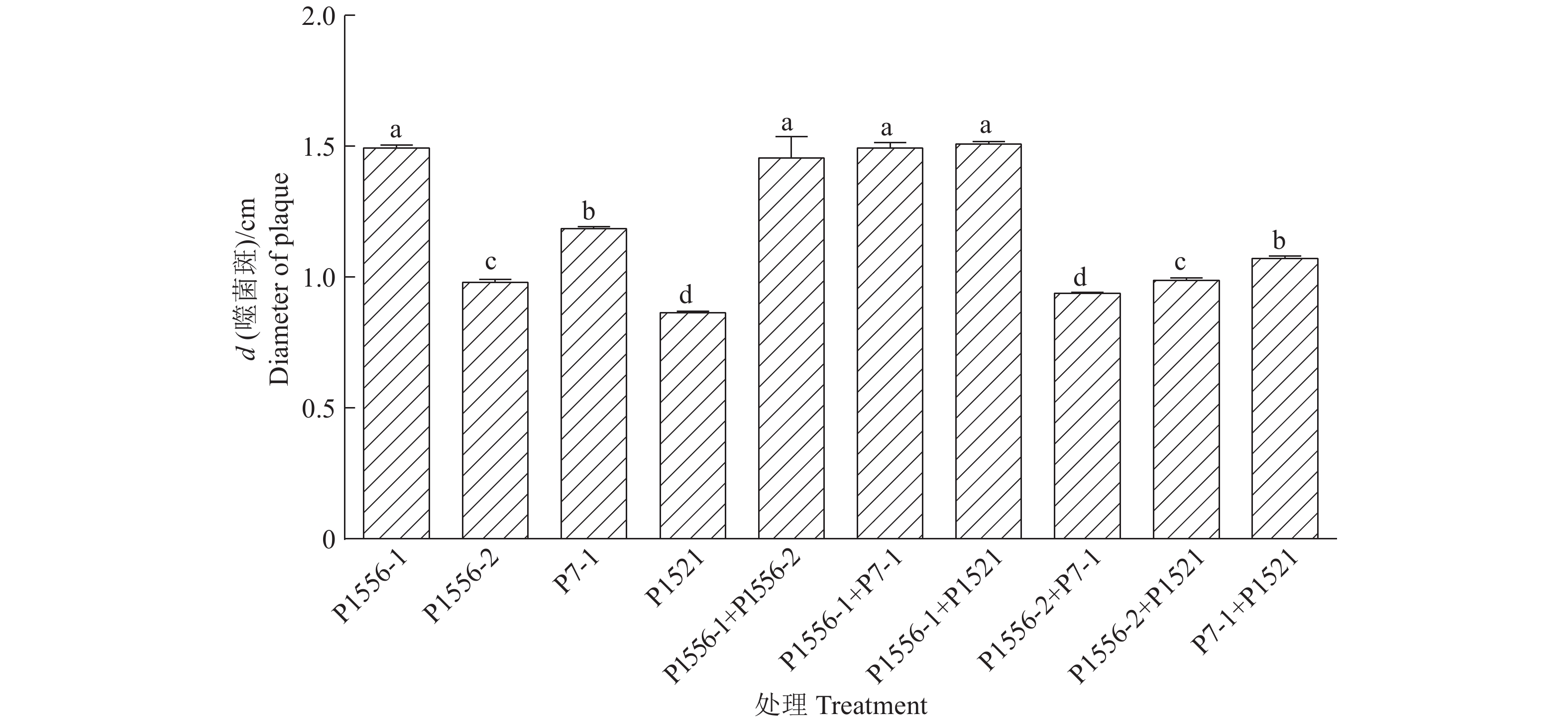
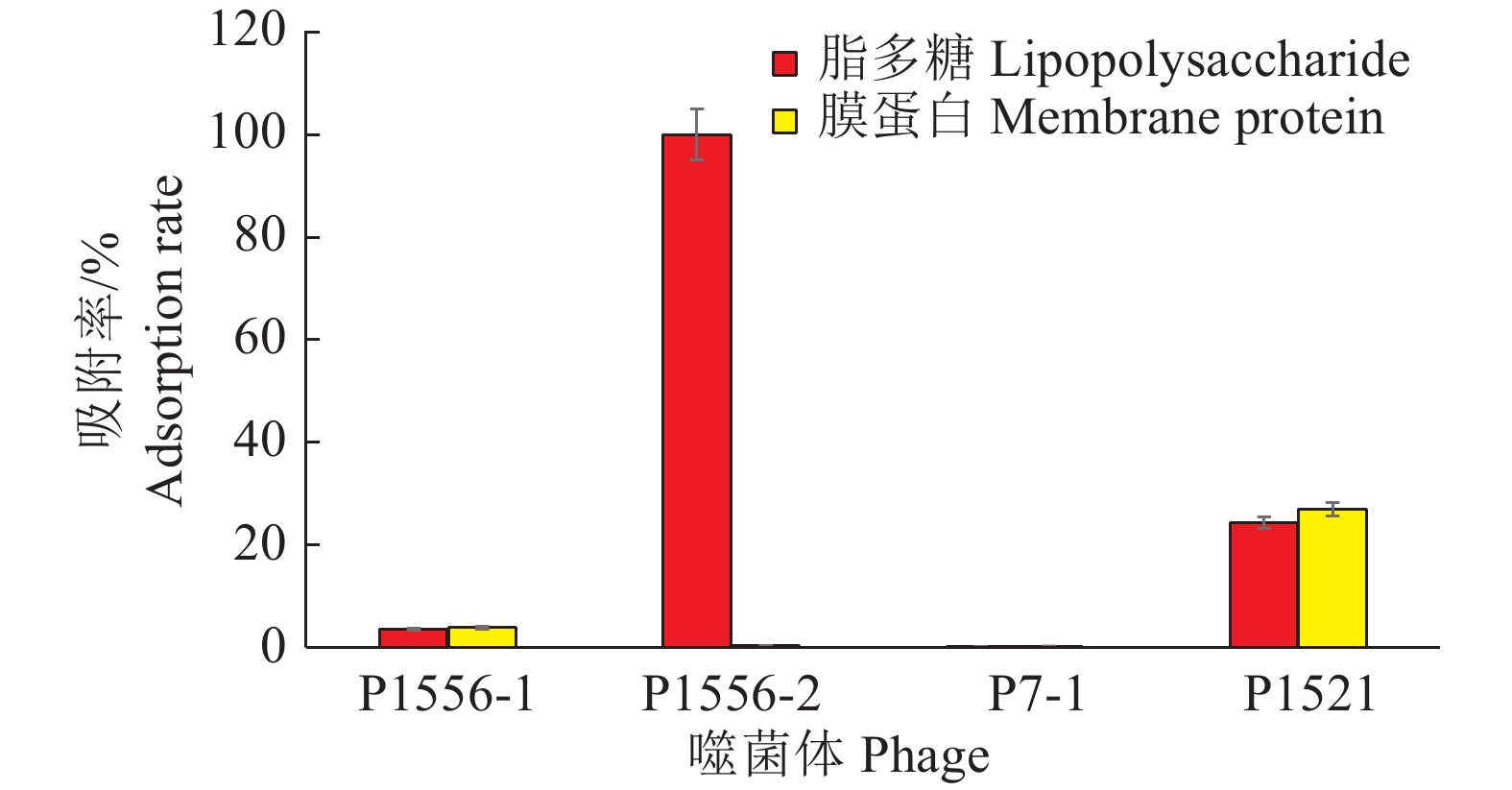
结果噬菌体P1556-1产生的噬菌斑最大,裂解青枯菌的能力强。4种噬菌体两两混合产生的噬菌斑与单一噬菌体产生的噬菌斑大小没有显著差异,但可延缓抗噬菌体的青枯菌产生。噬菌体P1556-2可被青枯菌的脂多糖吸附,P1521能被脂多糖和膜蛋白吸附,而P1556-1和P7-1均不能与脂多糖和膜蛋白作用。
结论不同噬菌体混合不能提高其裂解青枯菌的能力,但可以延缓抗性青枯菌的产生;不同噬菌体作用受体不同。
Abstract:ObjectiveTo investigate the lysis abilities of different phages mixtures against Ralstonia solanacearum, study their effects on the generation of phage-resistant strains, and clarify the receptors of four types of phages.
MethodFour types of phages, including P1556-1, P1556-2, P7-1 and P1521, were mixed in pairs. The sizes of plaques on R. solanacearum plates were determined. The resistance of R. solanacearum to phages was measured by co-culture of R. solanacearum and phages. Adsorption tests using phages, lipopolysaccharide (LPS) and membrane protein were conducted to determine the receptors of phages.
ResultPhage P1556-1 produced the largest plaque, indicating a strong lytic ability against R. solanacearum. The plaque sizes from the mixtures of two types of phages did not significantly differ from those of individual phages, while mixing two types of phages could slow down the generation of phage-resistant strains. Phage P1556-2 could be adsorbed by LPS of R. solanacearum, phage P1521 could be adsorbed by both LPS and membrane protein, while phage P1556-1 and P7-1 could be adsorbed by neither LPS nor membrane protein.
ConclusionMixing different phages can not improve the lysis ability against R. solanacearum, but can delay the generation of resistant host strain. The receptors of different phages are different.
-
Keywords:
- Ralstonia solanacearum /
- lytic phage /
- host resistance /
- receptor /
- plaque
-
-
-
[1] RAHMANI R, ZARRINI G, SHEIKHZADEH F, et al. Effective phages as green antimicrobial agents against antibiotic-resistant hospital Escherichia coli[J]. Jundishapur J Microbiol, 2015, 8(2): e17744.
[2] SARKER S A, SULTANA S, REUTELER G, et al. Oral phage therapy of acute bacterial diarrhea with two coliphage preparations: A randomized trial in children from Bangladesh[J]. Ebiomedicine, 2016, 4: 124-137.
[3] WALL S, ZHANG J, ROSTAGNO M, et al. Phage therapy to reduce preprocessing Salmonella infections in market-weight swine[J]. Appl Environ Microb, 2010, 76(1): 48-53. doi: 10.1128/AEM.00785-09
[4] WANG Y, BARTON M, ELLIOTT L, et al. Bacteriophage therapy for the control of Vibrio harveyi in greenlip abalone (Haliotis laevigata)[J]. Aquaculture, 2017, 473: 251-258. doi: 10.1016/j.aquaculture.2017.01.003
[5] SUSIANTO G, FARID M M, DHANY N R, et al. Host range for bacteriophages that infect bacterial blight pathogen on soybean[J]. Proc Environ Sci, 2014, 20: 760-766. doi: 10.1016/j.proenv.2014.03.091
[6] FLAHERTY J E, JONES J B, HARBAUGH B K, et al. Control of bacterial spot on tomato in the greenhouse and field with H-mutant bacteriophages[J]. Hortscience, 2000, 35(5): 882-884. doi: 10.21273/HORTSCI.35.5.882
[7] FLAHERTY J E, HARBAUGH B K, JONES J B, et al. H-mutant bacteriophages as a potential biocontrol of bacterial blight of geranium[J]. Hortscience, 2001, 36(1): 98-100. doi: 10.21273/HORTSCI.36.1.98
[8] BALOGH B, CANTEROS B I, STALL R E, et al. Control of citrus canker and citrus bacterial spot with bacteriophages[J]. Plant Dis, 2008, 92(7): 1048-1052. doi: 10.1094/PDIS-92-7-1048
[9] SCHNABEL E L, FERNANDO W G D, MEYER M P, et al. Bacteriophage of Erwinia amylovora and their potential for biocontrol[J]. Acta Hortic, 1999, 489(489): 649-654.
[10] RAVENSDALE M, BLOM T J, GRACIAl-GARZA J A, et al. Bacteriophages and the control of Erwinia carotovora subsp. carotovora[J]. Can J Plant Pathol, 2007, 29(2): 121-130. doi: 10.1080/07060660709507448
[11] ADDY H S, ASKORA A, KAWASAKI T, et al. Loss of virulence of the phytopathogen Ralstonia solanacearum through infection by φRSM filamentous phages[J]. Phytopathology, 2012, 102(5): 469. doi: 10.1094/PHYTO-11-11-0319-R
[12] ADDY H S, ASKORA A, KAWASAKI T, et al. Utilization of filamentous phage phi RSM3 to control bacterial wilt caused by Ralstonia solanacearum[J]. Plant Dis, 2012, 96(8): 1204-1209. doi: 10.1094/PDIS-12-11-1023-RE
[13] FUJIWARA A, FUJISAWA M, HAMASAKI R, et al. Biocontrol of Ralstonia solanacearum by treatment with lytic bacteriophages[J]. Appl Environ Microb, 2011, 77(12): 4155-4162.
[14] 康洁. 3种方法提取大肠杆菌E.coli脂多糖的比较[J]. 中国农学通报, 2010, 26(21): 12-15. [15] 余成鹏, 胡蓉花, 陈小强, 等. 江西和广东烟草青枯菌对噬菌体的敏感性及聚类分析[J]. 江西农业大学学报, 2018(4): 699-707. [16] WEI C H, LIU J L, MAINA A N, et al. Developing a bacteriophage cocktail for biocontrol of potato bacterial wilt[J]. Virol Sin, 2017, 32(6): 476-484.
[17] BAE J Y, WU J, LEE H J, et al. Biocontrol potential of a lytic bacteriophage PE204 against bacterial wilt of tomato[J]. J Microbiol Biotechn, 2012, 22(12): 1613-1620. doi: 10.4014/jmb.1208.08072
[18] MARRAFFINI L A. CRISPR-Cas immunity against phages: Its effects on the evolution and survival of bacterial pathogens[J]. PLoS Pathog, 2013, 9(12): e1003765. doi: 10.1371/journal.ppat.1003765
[19] BARRANGOU R, FREMAUX C, DEVEAU H, et al. CRISPR provides acquired resistance against viruses in prokaryotes[J]. Science, 2007, 315(5819): 1709. doi: 10.1126/science.1138140
[20] HYNES A P, VILLION M, MOINEAU S. Adaptation in bacterial CRISPR-Cas immunity can be driven by defective phages[J]. Nat Commun, 2014, 5: 4399. doi: 10.1038/ncomms5399
[21] WEI Y, CHESNE M T, TERNS R M, et al. Sequences spanning the leader-repeat junction mediate CRISPR adaptation to phage in Streptococcus thermophilus[J]. Nucleic Acids Res, 2015, 43(3): 1749-1758. doi: 10.1093/nar/gku1407
[22] SORENSEN M C H, van ALPHEN L B, HARBOE A, et al. Bacteriophage F336 recognizes the capsular phosphoramidate modification of Campylobacter jejuni NCTC11168[J]. J Bacteriol, 2011, 193(23): 6742-6749. doi: 10.1128/JB.05276-11
[23] ZALESKI P. The role of Dam methylation in phase variation of Haemophilus influenzae genes involved in defence against phage infection[J]. Microbiology, 2005, 151(10): 3361-3369. doi: 10.1099/mic.0.28184-0
[24] KIM M, RYU S. Characterization of a T5-Like coliphage, SPC35, and differential development of resistance to SPC35 in Salmonella enteric serovar Typhimurium and Escherichia coli[J]. Appl Environ Microb, 2011, 77(6): 2042-2050. doi: 10.1128/AEM.02504-10
[25] 毛普加, 洪愉, 毛小萍, 等. 噬菌体受体及其鉴定方法[J]. 中国微生态学杂志, 2014(6): 731-734. [26] BAE H, CHO Y. Complete genome sequence of Pseudomonas aeruginosa podophage MPK7, which requires type IV pili for infection[J]. Genome Announc, 2013, 1(5): e00744-13.
[27] NARULITA E, ADDY H S, KAWASAKI T, et al. The involvement of the PilQ secretin of type IV pili in phage infection in Ralstonia solanacearum[J]. Biochem Bioph Res Co, 2016, 469(4): 868-872. doi: 10.1016/j.bbrc.2015.12.071


 下载:
下载: