Abundance and structure diversity of denitrifying bacterial community in sediments of Guangzhou Liuxi River
-
摘要:目的
研究广州市流溪河沉积物中反硝化细菌的多样性,探讨环境因子对反硝化细菌群落结构和丰度的影响。
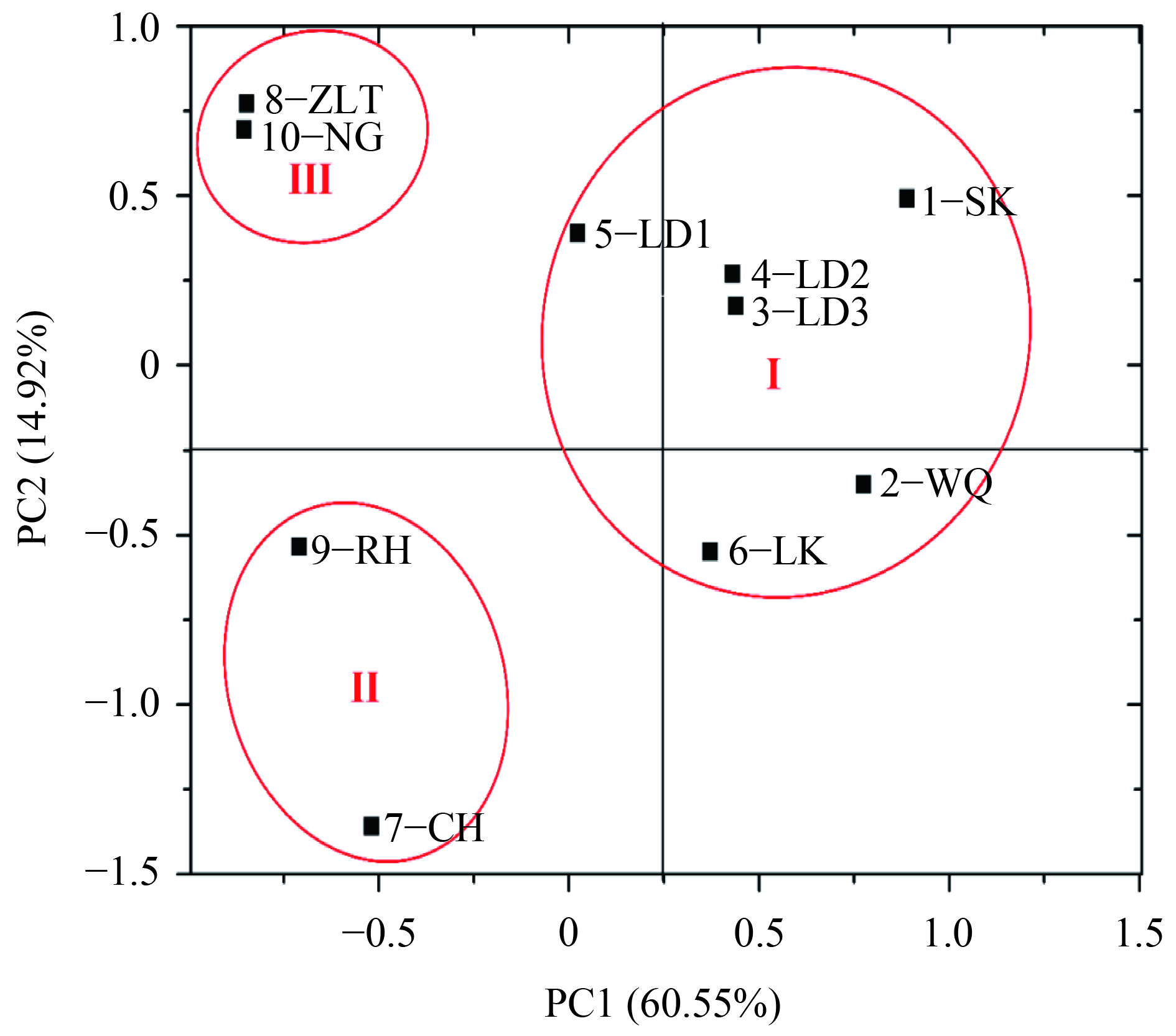
方法通过构建基因文库和实时荧光定量PCR(RT-qPCR)对沉积物中反硝化细菌的群落结构和丰度进行分析,并结合主坐标分析(PCoA)和冗余分析(RDA)研究反硝化细菌群落与环境因子之间的相关性。
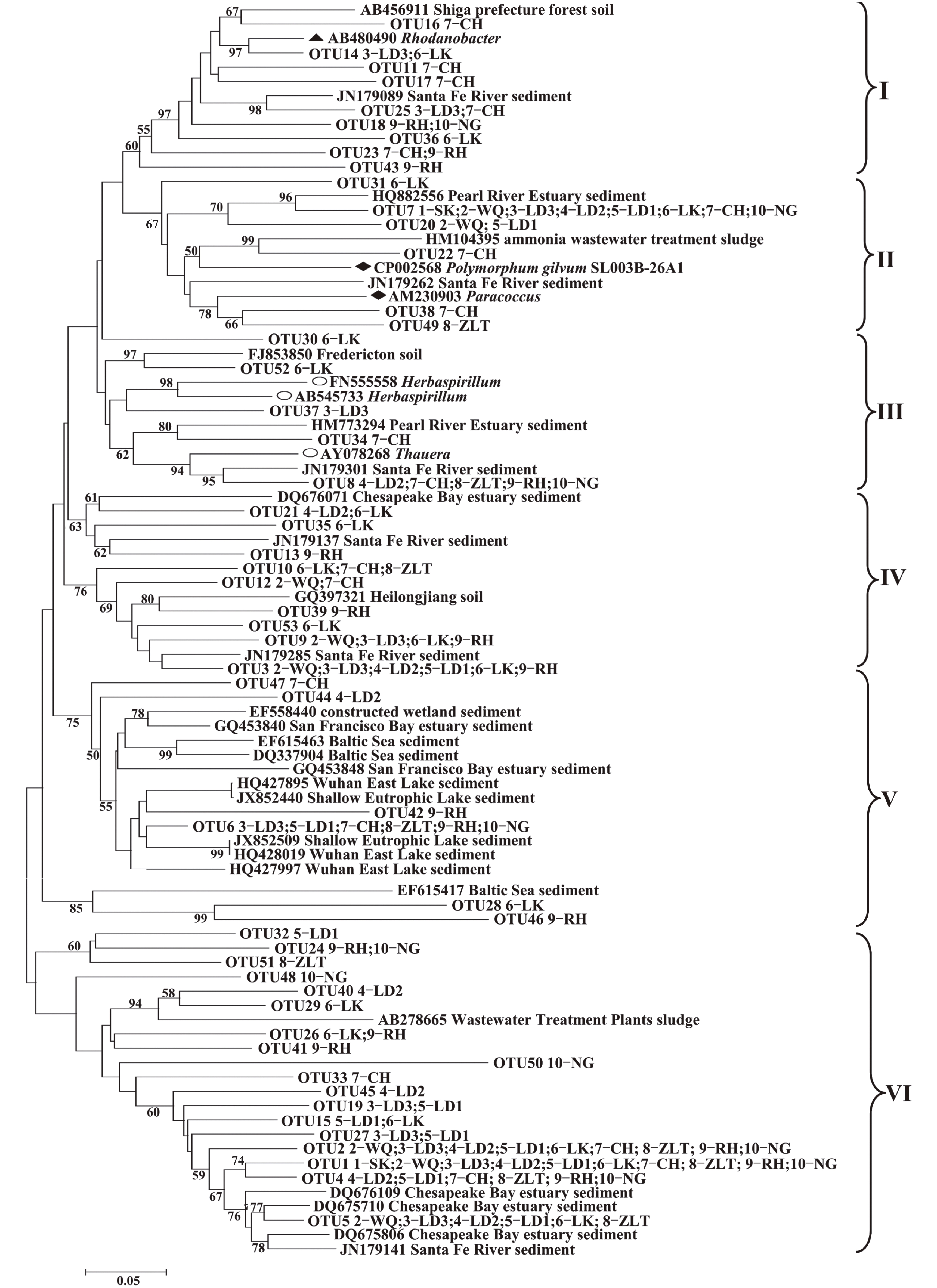
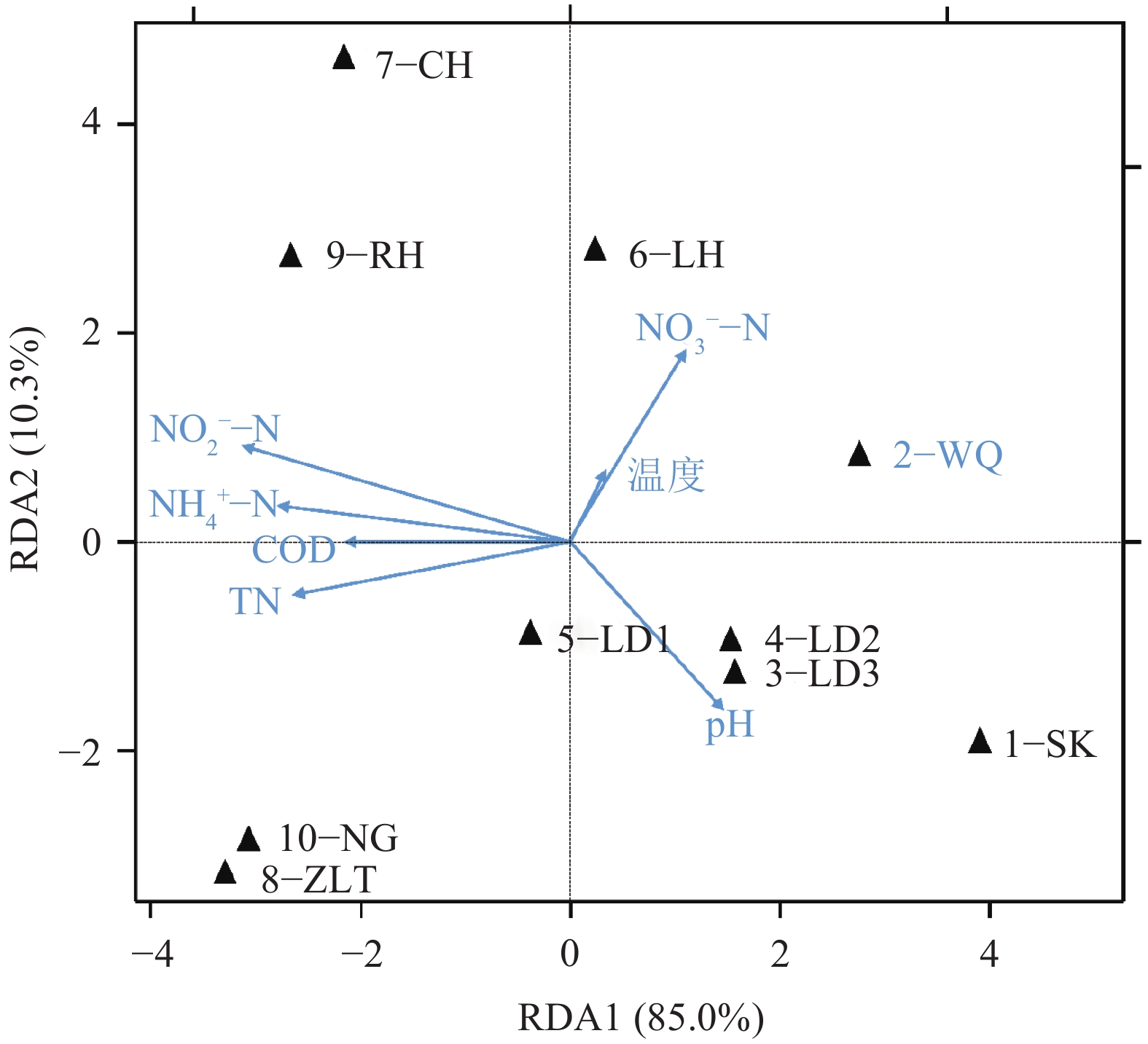
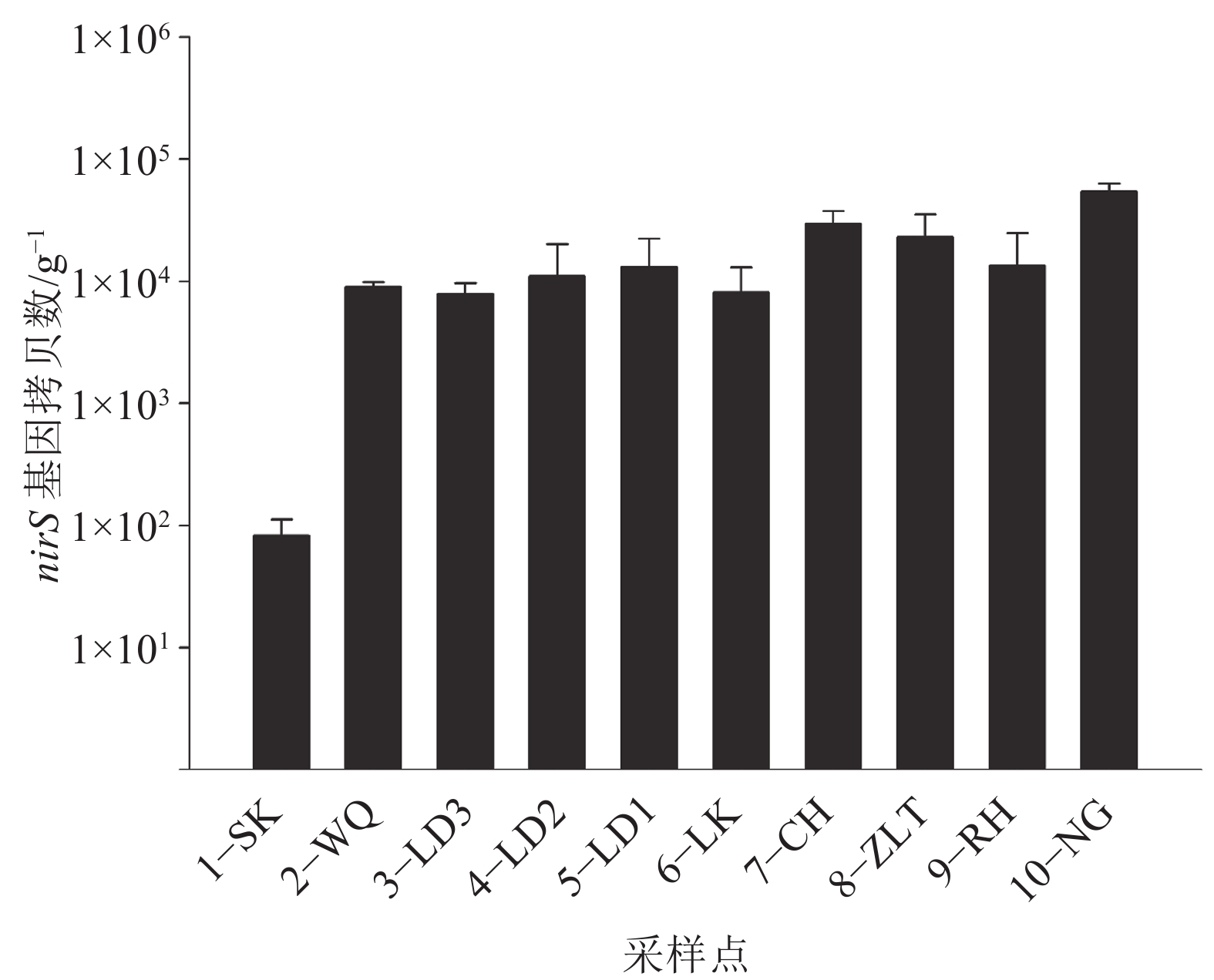
结果流溪河流域不同采样位点沉积物中nirS型反硝化细菌的组成和丰度存在明显差异。沉积物中大多数nirS序列与已知的反硝化细菌亲缘关系较远,而与其他环境样品中的微生物亲缘关系较近,与流溪河沉积物中nirS反硝化细菌亲缘关系较近的物种有产黄杆菌属Rhodanobacter、副球菌属Paracoccus、Polymorphum gilvum、草螺菌属Herbaspirillum和陶厄氏菌属Thauera。氨氮(NH4+–N)和硝酸盐(NO3––N)含量对反硝化细菌群落结构起决定性作用。反硝化细菌nirS基因拷贝数范围为8.26×101~5.45×104 g–1,nirS型反硝化细菌数量与化学需氧量(COD)、总氮(TN)、NH4+–N、和NO3––N含量之间存在显著相关性。
结论流溪河中化学污染物以及活性氮的大量存在显著影响了反硝化细菌群落结构和多样性。
Abstract:ObjectiveTo investigate the diversity of denitrifying bacteria in sediments of Guangzhou Liuxi River, and explore the effects of environmental factors on structure and abundance of bacterial community.
MethodThe community structure and abundance of denitrifying bacteria in sediments were analyzed by constructing gene library and real-time quantitative PCR (RT-qPCR). The principal coordinate analysis (PCoA) and redundancy analysis (RDA) were used to investigate the correlations between denitrifying bacterial community and environmental factors.
ResultThe composition and abundance of nirS-type denitrifier communities in different sampling sites of Liuxi River had distinctive discrepancy. Phylogenetic analysis showed that most of nirS gene sequences obtained in sediments had distant relationships with the known denitrifying bacteria, and the relationships with microorganisms from other environments were closer. The species that had close relationships with nirS-type denitrifier in sediments of Liuxi River were Rhodanobacter, Paracoccus, Polymorphum gilvum, Herbaspirillum and Thauera. Ammonium (NH4+–N) and nitrate (NO3––N) contents had decisive effects on the structure of denitrifying bacterial community. ThenirS gene copy numbers were ranged from 8.26×101 to 5.45×104 g–1, and nirS-type denitrifier abundance was significantly correlated with chemical oxygen demand, total nitrogen, ammonium and nitrate concentrations.
ConclusionThe abundant occurrence of chemical contaminants and reactive nitrogen in Liuxi River significantly influences denitrifying bacterial community structure and diversity.
-
Keywords:
- denitrifying bacteria /
- community structure /
- abundance /
- environmental factor /
- Liuxi River /
- sediment
-
-
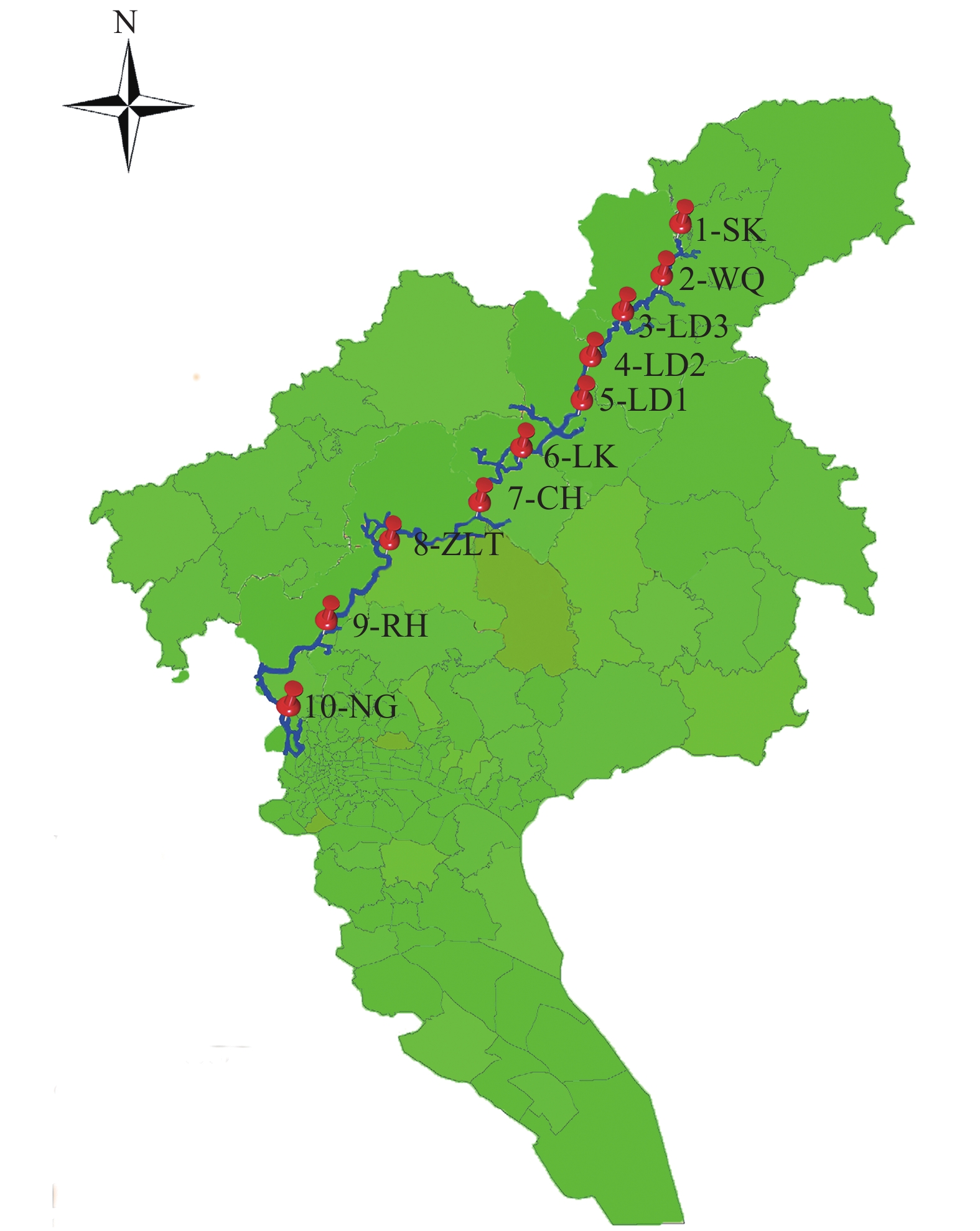
表 1 流溪河中各采样点底泥理化性质
Table 1 Physical and chemical properties of sediments in sampling sites within the Liuxi River
采样点 pH θ/℃ w/(mg·kg–1) COD NH4+–N NO2––N NO3––N TN 1–SK 7.66 29.6 6.67 0.33 0.08 3.11 4.39 2–WQ 7.51 29.8 21.33 6.09 0.02 3.21 9.36 3–LD3 7.59 28.7 21.33 1.34 0.08 2.79 4.71 4–LD2 7.35 27.7 21.33 6.45 0.03 3.53 10.04 5–LD1 7.48 29.5 10.67 4.79 0.06 2.83 9.50 6–LK 7.38 29.0 12.00 5.58 0.08 3.34 6.94 7–CH 7.33 29.0 41.33 18.45 0.29 2.93 30.94 8–ZLT 7.33 29.4 16.00 9.38 0.23 2.97 21.52 9–RH 7.49 29.5 29.33 10.98 0.26 3.67 25.32 10–NG 7.62 28.4 69.33 26.52 0.23 2.19 71.35 表 2 流溪河中不同采样位点nirS型反硝化细菌多样性指数
Table 2 Diversity characteristics of nirS-type denitrifiers in sampling sites in the Liuxi River
采样点 克隆子数量 OTU数量 Simpson指数 Shannon指数 Chao1 指数 Pielou指数 ACE指数 覆盖率/% 1–SK 52 2 0.92 0.16 2 0.23 2.0 100.0 2–WQ 48 8 0.59 1.11 11 0.53 13.6 91.6 3–LD3 51 12 0.35 1.50 17 0.60 65.5 86.3 4–LD2 50 11 0.37 1.48 16 0.62 33.1 88.0 5–LD1 52 12 0.17 1.95 33 0.78 52.6 86.5 6–LK 50 19 0.12 2.45 30 0.83 50.9 78.0 7–CH 51 19 0.07 2.68 22 0.91 25.8 86.3 8–ZLT 47 9 0.28 1.59 12 0.72 16.8 91.4 9–RH 45 17 0.08 2.48 62 0.87 47.8 77.8 10–NG 49 10 0.26 1.67 15 0.73 23.9 89.8 表 3 nirS基因丰度与环境因子相关系数
Table 3 Correlation analyses of nirS gene abundance with environmental factors
N=10 理化性质 P1) R pH 0.044 0.904 θ –0.367 0.297 w(NH4+–N) 0.965** 0.000 w(NO2––N) 0.609 0.061 w(NO3––N) –0.683* 0.029 w(COD) 0.948** 0.001 w(TN) 0.970** 0.000 1) “*” 、“**”分别表示在 0.05 和 0.01 水平相关性显著 -
[1] XIONG W, SUN Y, DING X, et al. Antibiotic resistance genes occurrence and bacterial community composition in the Liuxi River[J]. Front Environ Sci, 2014, 2. doi: 10.3389/fenvs.2014.00061
[2] CAFFREY J M, BANO N, KALANETRA K, et al. Ammonia oxidation and ammonia-oxidizing bacteria and archaea from estuaries with differing histories of hypoxia[J]. ISME J, 2007, 1(7): 660-662.
[3] ZUMFT W G. Cell Biology and molecular basis of denitrification[J]. Microbiol Mol Biol Rev, 1997, 61(4): 533-616.
[4] PHILIPPOT L, HØJBERG O. Dissimilatory nitrate reductases in bacteria[J]. Biochim Biophys Acta, 1999, 1446(1/2): 1-23.
[5] ABELL G C J, REVILL A T, SMITH C, et al. Archaeal ammonia oxidizers and nirS-type denitrifiers dominate sediment nitrifying and denitrifying populations in a subtropical macrotidal estuary[J]. ISME J, 2010, 4(2): 286-300.
[6] NOGALES B, TIMMIS K N, NEDWELL D B, et al. Detection and diversity of expressed denitrification genes in estuarine sediments after reverse transcription-PCR amplification from mRNA[J]. Appl Environ Microbiol, 2002, 68(10): 5017-5025.
[7] ZHENG Y, HOU L, LIU M, et al. Diversity, abundance, and distribution of nirS-harboring denitrifiers in intertidal sediments of the Yangtze Estuary[J]. Microb Ecol, 2015, 70(1): 30-40.
[8] DANG H, WANG C, LI J, et al. Diversity and distribution of sediment nirS-encoding bacterial assemblages in response to environmental gradients in the eutrophied Jiaozhou Bay, China[J]. Microb Ecol, 2009, 58(1): 161-169.
[9] THROBÄCK I N, ENWALL K, JARVIS Å, et al. Reassessing PCR primers targeting nirS, nirK and nosZ genes for community surveys of denitrifying bacteria with DGGE[J]. FEMS Microbiol Ecol, 2004, 49(3): 401-417.
[10] HENRY S, BRU D, STRES B, et al. Quantitative detection of the nosZ gene, encoding nitrous oxide reductase, and comparison of the abundances of 16S rRNA, narG, nirK, and nosZ genes in soils[J]. Appl Environ Microbiol, 2006, 72(8): 5181-5189.
[11] GAO J, HOU L, ZHENG Y, et al. nirS-Encoding denitrifier community composition, distribution, and abundance along the coastal wetlands of China[J]. Appl Microbiol Biotechnol, 2016, 100(19): 8573-8582.
[12] AVRAHAMI S, CONRAD R, BRAKER G. Effect of soil ammonium concentration on N2O release and on the community structure of ammonia oxidizers and denitrifiers[J]. Appl Environ Microbiol, 2002, 68(11): 5685-5692.
[13] FRANCIS C A, O'MULLAN G D, CORNWELL J C, et al. Transitions in nirS-type denitrifier diversity, community composition, and biogeochemical activity along the Chesapeake Bay estuary[J]. Front Microbiol, 2013, 4(237). doi: 10.3389/fmicb.2013.00237
[14] WANG Y, MA X, ZHOU S, et al. Expression of the nirS, hzsA, and hdh genes in response to nitrite shock and recovery in Candidatus kuenenia stuttgartiensis[J]. Environ Sci Technol, 2016, 50(13SI): 6940-6947.
[15] ROUSK J, BÅÅTH E, BROOKES P C, et al. Soil bacterial and fungal communities across a pH gradient in an arable soil[J]. ISME J, 2010, 4(10): 1340-1351.
[16] SZUKICS U, HACKL E, ZECHMEISTER-BOLTENSTERN S, et al. Rapid and dissimilar response of ammonia oxidizing archaea and bacteria to nitrogen and water amendment in two temperate forest soils[J]. Microbiol Res, 2012, 167(2): 103-109.
[17] 莫旭华, 麻威, 史荣久, 等. 氮肥对小麦田土壤nirS型反硝化细菌多样性的影响[J]. 微生物学报, 2009, 49(9): 1203-1208. [18] ZHANG Y, XIE X, JIAO N, et al. Diversity and distribution of amoA-type nitrifying and nirS-type denitrifying microbial communities in the Yangtze River estuary[J]. Biogeosciences, 2014, 11(8): 2131-2145.
[19] MOSIER A C, FRANCIS C A. Denitrifier abundance and activity across the San Francisco Bay estuary[J]. Environ Microbiol Rep, 2010, 2(5): 667-676.
[20] DONG L F, SMITH C J, PAPASPYROU S, et al. Changes in benthic denitrification, nitrate ammonification, and anammox process rates and nitrate and nitrite reductase gene abundances along an estuarine nutrient gradient (the Colne Estuary, United Kingdom)[J]. Appl Environ Microbiol, 2009, 75(10): 3171-3179.
[21] 胡玲珍, 陈振楼. 河口沉积物反硝化反应影响因子综述[J]. 环境科学动态, 2003(4): 41-43. -
期刊类型引用(2)
1. 张逸风,赵瑜君,苏平,吴晓毅,夏梦,高伟,黄璐琦. 比较蛋白质组学揭示茉莉酸甲酯诱导的雷公藤甲素生物合成. 中国现代中药. 2024(02): 316-327 .  百度学术
百度学术
2. 王婷,来欢欢,赵微,崔美林,张秀红. 基于蛋白质组学技术对植物乳杆菌YP36细菌素的抑菌机制研究. 中国酿造. 2024(04): 98-104 .  百度学术
百度学术
其他类型引用(3)


 下载:
下载: